The kinetically dominant assembly pathway for centrosomal asters in Caenorhabditis elegans is gamma-tubulin dependent
- PMID: 12011109
- PMCID: PMC2173857
- DOI: 10.1083/jcb.200202047
The kinetically dominant assembly pathway for centrosomal asters in Caenorhabditis elegans is gamma-tubulin dependent
Abstract
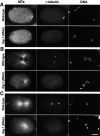
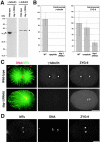
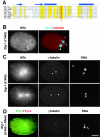
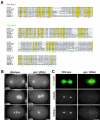
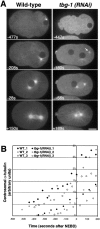
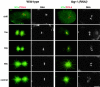
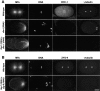
gamma-Tubulin-containing complexes are thought to nucleate and anchor centrosomal microtubules (MTs). Surprisingly, a recent study (Strome, S., J. Powers, M. Dunn, K. Reese, C.J. Malone, J. White, G. Seydoux, and W. Saxton. Mol. Biol. Cell. 12:1751-1764) showed that centrosomal asters form in Caenorhabditis elegans embryos depleted of gamma-tubulin by RNA-mediated interference (RNAi). Here, we investigate the nucleation and organization of centrosomal MT asters in C. elegans embryos severely compromised for gamma-tubulin function. We characterize embryos depleted of approximately 98% centrosomal gamma-tubulin by RNAi, embryos expressing a mutant form of gamma-tubulin, and embryos depleted of a gamma-tubulin-associated protein, CeGrip-1. In all cases, centrosomal asters fail to form during interphase but assemble as embryos enter mitosis. The formation of these mitotic asters does not require ZYG-9, a centrosomal MT-associated protein, or cytoplasmic dynein, a minus end-directed motor that contributes to self-organization of mitotic asters in other organisms. By kinetically monitoring MT regrowth from cold-treated mitotic centrosomes in vivo, we show that centrosomal nucleating activity is severely compromised by gamma-tubulin depletion. Thus, although unknown mechanisms can support partial assembly of mitotic centrosomal asters, gamma-tubulin is the kinetically dominant centrosomal MT nucleator.
Figures
References
-
- Alberts, B., D. Bray, J. Lewis, M. Raff, K. Roberts, and J. Watson. 1994. Molecular Biology of the Cell. Garland Publishing, Inc., New York. 1294 pp.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

