Replicational organization of three weakly expressed loci in Physarum polycephalum
- PMID: 12034812
- PMCID: PMC117180
- DOI: 10.1093/nar/30.11.2261
Replicational organization of three weakly expressed loci in Physarum polycephalum
Abstract
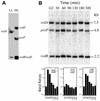
We previously mapped early-activated replication origins in the promoter regions of five abundantly transcribed genes in the slime mold Physarum polycephalum. This physical linkage between origins and genes is congruent with the preferential early replication of the active genes in mammalian cells. To determine how general this replicational organization is in the synchronous plasmodium of Physarum, we analyzed the replication of three weakly expressed genes. Bromodeoxyuridine (BrdUrd) density-shift and gene dosage experiments indicated that the redB (regulated in development) and redE genes replicate early, whereas redA replicates in mid-S phase. Bi-dimensional gel electrophoresis revealed that redA coincides with an origin that appears to be activated within a large temporal window in S phase so that the replication of the gene is not well defined temporally. The early replication of the redB and redE genes is due to the simultaneous activation of flanking origins at the onset of S phase. As a result, these two genes correspond to termination sites of DNA replication. Our data demonstrate that not all the Physarum promoters are preferred sites of initiation but, so far, all the expressed genes analyzed in detail either coincide with a replication origin or are embedded into a cluster of early firing replicons.
Figures
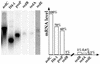



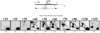
Similar articles
-
Low rate of replication fork progression lengthens the replication timing of a locus containing an early firing origin.Nucleic Acids Res. 2007;35(17):5763-74. doi: 10.1093/nar/gkm586. Epub 2007 Aug 23. Nucleic Acids Res. 2007. PMID: 17717000 Free PMC article.
-
Early activated replication origins within the cell cycle-regulated histone H4 genes in Physarum.Nucleic Acids Res. 1999 May 15;27(10):2091-8. doi: 10.1093/nar/27.10.2091. Nucleic Acids Res. 1999. PMID: 10219081 Free PMC article.
-
Mapping of a replication origin within the promoter region of two unlinked, abundantly transcribed actin genes of Physarum polycephalum.Mol Cell Biol. 1996 Mar;16(3):968-76. doi: 10.1128/MCB.16.3.968. Mol Cell Biol. 1996. PMID: 8622700 Free PMC article.
-
Maternal inheritance of mitochondria: multipolarity, multiallelism and hierarchical transmission of mitochondrial DNA in the true slime mold Physarum polycephalum.J Plant Res. 2010 Mar;123(2):139-48. doi: 10.1007/s10265-009-0298-5. J Plant Res. 2010. PMID: 20082112 Review.
-
Genetic Diversity in the mtDNA of Physarum polycephalum.Genes (Basel). 2023 Mar 2;14(3):628. doi: 10.3390/genes14030628. Genes (Basel). 2023. PMID: 36980901 Free PMC article. Review.
Cited by
-
Developmentally regulated usage of Physarum DNA replication origins.EMBO Rep. 2003 May;4(5):474-8. doi: 10.1038/sj.embor.embor822. EMBO Rep. 2003. PMID: 12776736 Free PMC article.
-
Low rate of replication fork progression lengthens the replication timing of a locus containing an early firing origin.Nucleic Acids Res. 2007;35(17):5763-74. doi: 10.1093/nar/gkm586. Epub 2007 Aug 23. Nucleic Acids Res. 2007. PMID: 17717000 Free PMC article.
-
RNase-dependent discontinuities associated with the crossovers of spontaneously formed joint DNA molecules in Physarum polycephalum.Chromosoma. 2010 Dec;119(6):601-11. doi: 10.1007/s00412-010-0281-x. Epub 2010 Jul 7. Chromosoma. 2010. PMID: 20607271
References
-
- Simon I. and Cedar,H. (1996) Temporal order of DNA replication. In DePamphilis,M.L. (ed.), DNA Replication in Eukaryotic Cells. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp. 387–408.
-
- Wohlschlegel J.A., Dwyer,B.T., Dhar,S.K., Cvetic,C., Walter,J.C. and Dutta,A. (2000) Inhibition of eukaryotic DNA replication by geminin binding to Cdt1. Science, 290, 2309–2312. - PubMed
-
- Nguyen V.Q., Co,C. and Li,J.J. (2001) Cyclin-dependent kinases prevent DNA re-replication through multiple mechanisms. Nature, 411, 1068–1073. - PubMed
-
- Diffley J.F. (2001) DNA replication: building the perfect switch. Curr. Biol., 11, R367–R370. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

