A major susceptibility locus for specific language impairment is located on 13q21
- PMID: 12048648
- PMCID: PMC384992
- DOI: 10.1086/341095
A major susceptibility locus for specific language impairment is located on 13q21
Abstract
Children who fail to develop language normally-in the absence of explanatory factors such as neurological disorders, hearing impairment, or lack of adequate opportunity-are clinically described as having specific language impairment (SLI). SLI has a prevalence of approximately 7% in children entering school and is associated with later difficulties in learning to read. Research indicates that genetic factors are important in the etiology of SLI. Studies have consistently demonstrated that SLI aggregates in families. Increased monozygotic versus dizygotic twin concordance rates indicate that heredity, not just shared environment, is the cause of the familial clustering. We have collected five pedigrees of Celtic ancestry that segregate SLI, and we have conducted genomewide categorical linkage analysis, using model-based LOD score techniques. Analysis was conducted under both dominant and recessive models by use of three phenotypic classifications: clinical diagnosis, language impairment (spoken language quotient <85) and reading discrepancy (nonverbal IQ minus non-word reading >15). Chromosome 13 yielded a maximum multipoint LOD score of 3.92 under the recessive reading discrepancy model. Simulation to correct for multiple models and multiple phenotypes indicated that the genomewide empirical P value is <.01. As an alternative measure, we also computed the posterior probability of linkage (PPL), obtaining a PPL of 53% in the same region. One other genomic region yielded suggestive results on chromosome 2 (multipoint LOD score 2.86, genomic P value <.06 under the recessive language impairment model). Our findings underscore the utility of traditional LOD-score-based methods in finding genes for complex diseases, specifically, SLI.
Figures
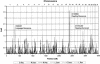



References
Electronic-Database Information
-
- CIDR, http://www.cidr.jhmi.edu/ (for genotyping protocols)
-
- Genome Database, http://www.gdb.org/ (for initial STS sequences used to redesign primers)
-
- Human Genome Project Working Draft, http://genome.ucsc.edu
-
- Marshfield Medical Center for Medical Genetics, http://research.marshfieldclinic.org/genetics/ (for order and genetic distances of microsatellite markers)
-
- National Center for Biotechnology Information (NCBI) Genetic Analysis Software, ftp://fastlink.nih.gov/pub/fastlink/ (for FASTLINK versions of MLINK and LINKMAP)
References
-
- Aram DM (1991) Comments on specific language impairment as a clinical category. Lang Speech Hear Serv School 22:84–87
-
- Benasich AA, Spitz RV (1998) Insights from infants: temporal processing abilities and genetics contribute to language development. In: Willems G, Whitmore K (eds) A neurodevelopmental approach to specific learning disorders. MacKeith Press, London, pp 191–210
-
- Benasich AA, Tallal P (1996) Auditory temporal processing thresholds, habituation, and recognition memory over the first year. Infant Behav Dev 19:339–357
-
- Bishop DVM (1994) Is specific language impairment a valid diagnostic category? Genetic and psycholinguistic evidence. Philos Trans R Soc Lond B Biol Sci 346:105–111 - PubMed
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

