Two-component signal transduction pathways in Arabidopsis
- PMID: 12068096
- PMCID: PMC161668
- DOI: 10.1104/pp.005504
Two-component signal transduction pathways in Arabidopsis
Abstract
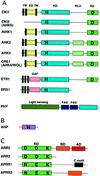
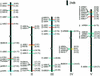
The two-component system, consisting of a histidine (His) protein kinase that senses a signal input and a response regulator that mediates the output, is an ancient and evolutionarily conserved signaling mechanism in prokaryotes and eukaryotes. The identification of 54 His protein kinases, His-containing phosphotransfer proteins, response regulators, and related proteins in Arabidopsis suggests an important role of two-component phosphorelay in plant signal transduction. Recent studies indicate that two-component elements are involved in plant hormone, stress, and light signaling. In this review, we present a genome analysis of the Arabidopsis two-component elements and summarize the major advances in our understanding of Arabidopsis two-component signaling.
Figures
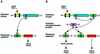




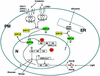
References
-
- AGI. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2000;408:796–815. - PubMed
-
- Bleecker AB, Kende H. Ethylene: a gaseous signal molecule in plants. Annu Rev Cell Dev Biol. 2000;16:1–18. - PubMed
-
- Chang C, Kwok SF, Bleecker AB, Meyerowitz EM. Arabidopsis ethylene-response gene ETR1: similarity of product to two-component regulators. Science. 1993;262:539–544. - PubMed
-
- Chang C, Stadler R. Ethylene hormone receptor action in Arabidopsis. Bioessays. 2001;23:619–627. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

