The pgp1 mutant locus of Arabidopsis encodes a phosphatidylglycerolphosphate synthase with impaired activity
- PMID: 12068104
- PMCID: PMC161686
- DOI: 10.1104/pp.002725
The pgp1 mutant locus of Arabidopsis encodes a phosphatidylglycerolphosphate synthase with impaired activity
Abstract
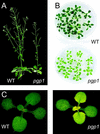
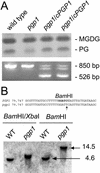
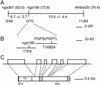
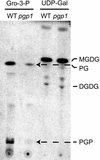
Phosphatidylglycerol is a ubiquitous phospholipid that is also present in the photosynthetic membranes of plants. Multiple independent lines of evidence suggest that this lipid plays a critical role for the proper function of photosynthetic membranes and cold acclimation. In eukaryotes, different subcellular compartments are competent for the biosynthesis of phosphatidylglycerol. Details on the plant-specific pathways in different organelles are scarce. Here, we describe a phosphatidylglycerol biosynthesis-deficient mutant of Arabidopsis, pgp1. The overall content of phosphatidylglycerol is reduced by 30%. This mutant carries a point mutation in the CDP-alcohol phosphotransferase motif of the phosphatidylglycerolphosphate synthase (EC 2.7.8.5) isoform encoded by a gene on chromosome 2. The mutant shows an 80% reduction in plastidic phosphatidylglycerolphosphate synthase activity consistent with the plastidic location of this particular isoform. Mutant plants are pale green, and their photosynthesis is impaired. This mutant provides a promising new tool to elucidate the biosynthesis and function of plastidic phosphatidylglycerol in seed plants.
Figures

References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2001;408:796–815. - PubMed
-
- Bechtold N, Pelletier G. In planta Agrobacterium-mediated transformation of adult Arabidopsis thalianaplants by vacuum infiltration. In: Salinas J, Martinez-Zapater JM, editors. Arabidopsis Protocols, Methods in Molecular Biology. Vol. 82. Totowa, NJ: Humana Press; 1998. pp. 259–266. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

