Complementary roles of multiple nuclear targeting signals in the capsid proteins of the parvovirus minute virus of mice during assembly and onset of infection
- PMID: 12072505
- PMCID: PMC136310
- DOI: 10.1128/jvi.76.14.7049-7059.2002
Complementary roles of multiple nuclear targeting signals in the capsid proteins of the parvovirus minute virus of mice during assembly and onset of infection
Abstract
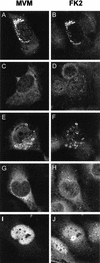
This report describes the distribution of conventional nuclear localization sequences (NLS) and of a beta-stranded so-called nuclear localization motif (NLM) in the two proteins (VP1, 82 kDa; VP2, 63 kDa) forming the T=1 icosahedral capsid of the parvovirus minute virus of mice (MVM) and their functions in viral biogenesis and the onset of infection. The approximately 10 VP1 molecules assembled in the MVM particle harbor in its 142-amino-acid (aa) N-terminal-specific region four clusters of basic amino acids, here called BC1 (aa 6 to 10), BC2 (aa 87 to 90), BC3 (aa 109 to 115), and BC4 (aa 126 to 130), that fit consensus NLS and an NLM placed toward the opposite end of the polypeptide (aa 670 to 680) found to be necessary for VP2 nuclear uptake. Deletions and site-directed mutations constructed in an infectious MVM plasmid showed that BC1, BC2, and NLM are cooperative nuclear transport sequences in singly expressed VP1 subunits and that they conferred nuclear targeting competence on the VP1/VP2 oligomers arising in normal infection, while BC3 and BC4 did not display nuclear transport activity. Notably, VP1 proteins mutated at BC1 and -2, and particularly with BC1 to -4 sequences deleted, induced nuclear and cytoplasmic foci of colocalizing conjugated ubiquitin that could be rescued from the ubiquitin-proteasome degradation pathway by the coexpression of VP2 and NS2 isoforms. These results suggest a role for VP2 in viral morphogenesis by assisting cytoplasmic folding of VP1/VP2 subviral complexes, which is further supported by the capacity of NLM-bearing transport-competent VP2 subunits to recruit VP1 into the nuclear capsid assembly pathway regardless of the BC composition. Instead, all four BC sequences, which are located in the interior of the capsid, were absolutely required by the incoming infectious MVM particle for the onset of infection, suggesting either an important conformational change or a disassembly of the coat for nuclear entry of a VP1-associated viral genome. Therefore, the evolutionarily conserved BC sequences and NLM domains provide complementary nuclear transport functions to distinct supramolecular complexes of capsid proteins during the autonomous parvovirus life cycle.
Figures




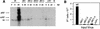
Similar articles
-
Nuclear transport of trimeric assembly intermediates exerts a morphogenetic control on the icosahedral parvovirus capsid.J Mol Biol. 2006 Mar 31;357(3):1026-38. doi: 10.1016/j.jmb.2006.01.019. Epub 2006 Jan 23. J Mol Biol. 2006. PMID: 16469332
-
A beta-stranded motif drives capsid protein oligomers of the parvovirus minute virus of mice into the nucleus for viral assembly.J Virol. 2000 Apr;74(8):3804-14. doi: 10.1128/jvi.74.8.3804-3814.2000. J Virol. 2000. PMID: 10729155 Free PMC article.
-
Low pH-dependent endosomal processing of the incoming parvovirus minute virus of mice virion leads to externalization of the VP1 N-terminal sequence (N-VP1), N-VP2 cleavage, and uncoating of the full-length genome.J Virol. 2006 Jan;80(2):1015-24. doi: 10.1128/JVI.80.2.1015-1024.2006. J Virol. 2006. PMID: 16379002 Free PMC article.
-
The role of nuclear localization signal in parvovirus life cycle.Virol J. 2017 Apr 14;14(1):80. doi: 10.1186/s12985-017-0745-1. Virol J. 2017. PMID: 28410597 Free PMC article. Review.
-
Role of capsid proteins in parvoviruses infection.Virol J. 2015 Aug 4;12:114. doi: 10.1186/s12985-015-0344-y. Virol J. 2015. PMID: 26239432 Free PMC article. Review.
Cited by
-
Nuclear export of the nonenveloped parvovirus virion is directed by an unordered protein signal exposed on the capsid surface.J Virol. 2004 Oct;78(19):10685-94. doi: 10.1128/JVI.78.19.10685-10694.2004. J Virol. 2004. PMID: 15367635 Free PMC article.
-
Minute virus of mice, a parvovirus, in complex with the Fab fragment of a neutralizing monoclonal antibody.J Virol. 2007 Sep;81(18):9851-8. doi: 10.1128/JVI.00775-07. Epub 2007 Jul 11. J Virol. 2007. PMID: 17626084 Free PMC article.
-
Cytoplasmic Parvovirus Capsids Recruit Importin Beta for Nuclear Delivery.J Virol. 2020 Jan 31;94(4):e01532-19. doi: 10.1128/JVI.01532-19. Print 2020 Jan 31. J Virol. 2020. PMID: 31748386 Free PMC article.
-
Adeno-associated virus type 2 capsids with externalized VP1/VP2 trafficking domains are generated prior to passage through the cytoplasm and are maintained until uncoating occurs in the nucleus.J Virol. 2006 Nov;80(22):11040-54. doi: 10.1128/JVI.01056-06. Epub 2006 Sep 6. J Virol. 2006. PMID: 16956943 Free PMC article.
-
Protoparvovirus Cell Entry.Viruses. 2017 Oct 26;9(11):313. doi: 10.3390/v9110313. Viruses. 2017. PMID: 29072600 Free PMC article. Review.
References
-
- Agbandje-McKenna, M., A. Llamas-Saiz, F. Wang, P. Tattersall, and M. G. Rossmann. 1998. Functional implications of the structure of the murine parvovirus minute virus of mice. Structure 6:1369-1381. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

