Further characterization of equine foamy virus reveals unusual features among the foamy viruses
- PMID: 12072521
- PMCID: PMC136322
- DOI: 10.1128/jvi.76.14.7220-7227.2002
Further characterization of equine foamy virus reveals unusual features among the foamy viruses
Abstract
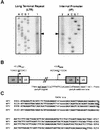
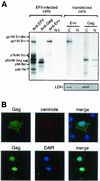
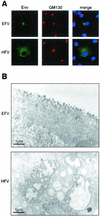
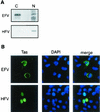
Foamy viruses (FVs) are nonpathogenic, widely spread complex retroviruses which have been isolated in nonhuman primates, cattle, cats, and more recently in horses. The equine foamy virus (EFV) was isolated from healthy horses and was characterized by molecular cloning and nucleotide sequence analysis. Here, to further characterize this new FV isolate, the location of the transcriptional cap and poly(A) addition sites as well as the main splice donor and acceptor sites were determined, demonstrating the existence of the specific subgenomic pol mRNA, one specific feature of FVs. Moreover, similar to what has been described for the human foamy virus (HFV), the prototype of FVs, a replication-defective EFV genome was identified during persistent infection. At the protein level, the use of specific antibodies allowed us to determine the size and the subcellular localization of EFV Gag, Env, and Tas, the viral transactivators. While EFV Gag was detected in both the cytoplasm and the nucleus, EFV Env mainly localized in the Golgi complex, in contrast to HFV Env, which is sequestered in the endoplasmic reticulum. In addition, electron microscopy analysis demonstrated that EFV budding occurs at the plasma membrane and not intracellularly, as is the case for primate FVs. Interestingly, EFV Tas was detected both in the nucleus and the cytoplasm of Tas-transfected cells, in contrast to the strict nuclear localization of other FV Tas but similar to the equine infectious anemia virus Tat gene product. Taken together, our results reveal that this new FV isolate exhibits remarkable features among FVs, bringing new insights into the biology of these unconventional retroviruses.
Figures


Similar articles
-
Isolation and characterization of an equine foamy virus.J Virol. 2000 May;74(9):4064-73. doi: 10.1128/jvi.74.9.4064-4073.2000. J Virol. 2000. PMID: 10756018 Free PMC article.
-
Characterization of the genome of feline foamy virus and its proteins shows distinct features different from those of primate spumaviruses.J Virol. 1997 Sep;71(9):6727-41. doi: 10.1128/JVI.71.9.6727-6741.1997. J Virol. 1997. PMID: 9261397 Free PMC article.
-
Characterization of the spliced pol transcript of feline foamy virus: the splice acceptor site of the pol transcript is located in gag of foamy viruses.J Virol. 1996 Dec;70(12):9024-7. doi: 10.1128/JVI.70.12.9024-9027.1996. J Virol. 1996. PMID: 8971036 Free PMC article.
-
Non-primate foamy viruses.Curr Top Microbiol Immunol. 2003;277:197-211. doi: 10.1007/978-3-642-55701-9_9. Curr Top Microbiol Immunol. 2003. PMID: 12908774 Review.
-
Regulation of foamy viral transcription and RNA export.Adv Virus Res. 2011;81:1-31. doi: 10.1016/B978-0-12-385885-6.00006-7. Adv Virus Res. 2011. PMID: 22094077 Review.
Cited by
-
Analysis of bovine foamy virus btas mRNA transcripts during persistent infection.Virus Genes. 2010 Feb;40(1):84-93. doi: 10.1007/s11262-009-0422-6. Epub 2009 Nov 13. Virus Genes. 2010. PMID: 19911263
-
Efficient production of inhibitor-free foamy virus glycoprotein-containing retroviral vectors by proteoglycan-deficient packaging cells.Mol Ther Methods Clin Dev. 2022 Aug 1;26:394-412. doi: 10.1016/j.omtm.2022.07.004. eCollection 2022 Sep 8. Mol Ther Methods Clin Dev. 2022. PMID: 36034773 Free PMC article.
-
Bovine foamy virus transactivator BTas interacts with cellular RelB to enhance viral transcription.J Virol. 2010 Nov;84(22):11888-97. doi: 10.1128/JVI.01036-10. Epub 2010 Sep 15. J Virol. 2010. PMID: 20844054 Free PMC article.
-
Investigating the intercellular spreading properties of the foamy virus Gag protein.PLoS One. 2012;7(2):e31108. doi: 10.1371/journal.pone.0031108. Epub 2012 Feb 29. PLoS One. 2012. PMID: 22393357 Free PMC article.
-
Non-simian foamy viruses: molecular virology, tropism and prevalence and zoonotic/interspecies transmission.Viruses. 2013 Sep 13;5(9):2169-209. doi: 10.3390/v5092169. Viruses. 2013. PMID: 24064793 Free PMC article. Review.
References
-
- Bock, M., M. Heinkelein, D. Lindemann, and A. Rethwilm. 1998. Cells expressing the human foamy virus (HFV) accessory Bet protein are resistant to productive HFV superinfection. Virology 250:194-204. - PubMed
-
- Bodem, J., M. Zemba, and R. M. Flügel. 1998. Nuclear localization of the functional Bel 1 transactivator but not of the gag proteins of the feline foamy virus. Virology 251:22-27. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

