A Salmonella enterica serovar typhimurium hemA mutant is highly susceptible to oxidative DNA damage
- PMID: 12081946
- PMCID: PMC135181
- DOI: 10.1128/JB.184.14.3774-3784.2002
A Salmonella enterica serovar typhimurium hemA mutant is highly susceptible to oxidative DNA damage
Abstract
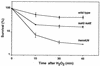
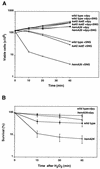
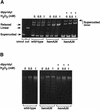
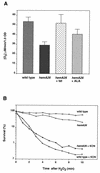
The first committed step in the biosynthesis of heme, an important cofactor of two catalases and a number of cytochromes, is catalyzed by the hemA gene product. Salmonella enterica serovar Typhimurium hemA26::Tn10d (hemA26) was identified in a genetic screen of insertion mutants that were sensitive to hydrogen peroxide. Here we show that the hemA26 mutant respires at half the rate of wild-type cells and is highly susceptible to the effects of oxygen species. Exposure of the hemA26 strain to hydrogen peroxide results in extensive DNA damage and cell death. The chelation of intracellular free iron fully abrogates the sensitivity of this mutant, indicating that the DNA damage results from the iron-catalyzed formation of hydroxyl radicals. The inactivation of heme synthesis does not change the amount of intracellular iron, but by diminishing the rate of respiration, it apparently increases the amount of reducing equivalents available to drive the Fenton reaction. We also report that hydrogen peroxide has opposite effects on the expression of hemA and hemH, the first and last genes of heme biosynthesis pathway, respectively. hemA mRNA levels decrease, while the transcription of hemH is induced by hydrogen peroxide, in an oxyR-dependent manner. The oxyR-dependent induction is suppressed under conditions that accelerate the Fenton reaction by a mechanism that is not yet understood.
Figures



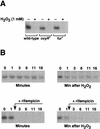

Similar articles
-
Fusidic acid-resistant mutants of Salmonella enterica serovar typhimurium have low levels of heme and a reduced rate of respiration and are sensitive to oxidative stress.Antimicrob Agents Chemother. 2004 Oct;48(10):3877-83. doi: 10.1128/AAC.48.10.3877-3883.2004. Antimicrob Agents Chemother. 2004. PMID: 15388448 Free PMC article.
-
Regulation of heme biosynthesis in Salmonella typhimurium: activity of glutamyl-tRNA reductase (HemA) is greatly elevated during heme limitation by a mechanism which increases abundance of the protein.J Bacteriol. 1997 May;179(9):2907-14. doi: 10.1128/jb.179.9.2907-2914.1997. J Bacteriol. 1997. PMID: 9139907 Free PMC article.
-
Transcription of the glutamyl-tRNA reductase (hemA) gene in Salmonella typhimurium and Escherichia coli: role of the hemA P1 promoter and the arcA gene product.J Bacteriol. 1996 Feb;178(3):638-46. doi: 10.1128/jb.178.3.638-646.1996. J Bacteriol. 1996. PMID: 8550494 Free PMC article.
-
The OxyR regulon.Antonie Van Leeuwenhoek. 1990 Oct;58(3):157-61. doi: 10.1007/BF00548927. Antonie Van Leeuwenhoek. 1990. PMID: 2256675 Review.
-
OxyR: a regulator of antioxidant genes.J Nutr. 1992 Mar;122(3 Suppl):627-30. doi: 10.1093/jn/122.suppl_3.627. J Nutr. 1992. PMID: 1542022 Review.
Cited by
-
Global transcriptome analysis of Staphylococcus aureus response to hydrogen peroxide.J Bacteriol. 2006 Feb;188(4):1648-59. doi: 10.1128/JB.188.4.1648-1659.2006. J Bacteriol. 2006. PMID: 16452450 Free PMC article.
-
Fusidic acid-resistant mutants of Salmonella enterica serovar typhimurium have low levels of heme and a reduced rate of respiration and are sensitive to oxidative stress.Antimicrob Agents Chemother. 2004 Oct;48(10):3877-83. doi: 10.1128/AAC.48.10.3877-3883.2004. Antimicrob Agents Chemother. 2004. PMID: 15388448 Free PMC article.
-
Altered linkage of hydroxyacyl chains in lipid A of Campylobacter jejuni reduces TLR4 activation and antimicrobial resistance.J Biol Chem. 2010 May 21;285(21):15828-36. doi: 10.1074/jbc.M110.102061. Epub 2010 Mar 29. J Biol Chem. 2010. PMID: 20351099 Free PMC article.
-
Heme-dependent metalloregulation by the iron response regulator (Irr) protein in Rhizobium and other Alpha-proteobacteria.Biometals. 2009 Feb;22(1):89-97. doi: 10.1007/s10534-008-9192-1. Epub 2008 Dec 18. Biometals. 2009. PMID: 19093075 Free PMC article.
-
Characterization of enteropathogenic Escherichia coli mutants that fail to disrupt host cell spreading and attachment to substratum.Infect Immun. 2006 Feb;74(2):839-49. doi: 10.1128/IAI.74.2.839-849.2006. Infect Immun. 2006. PMID: 16428726 Free PMC article.
References
-
- Andrews, S. C. 1998. Iron storage in bacteria. Adv. Microb. Physiol. 40:281-351. - PubMed
-
- Beale, S. I. 1996. Biosynthesis of hemes, p. 731-748. In F. C. Neidhardt, R. Curtiss III, J. L. Ingraham, E. C. C. Lin, K. B. Low, B. Magasanik, W. S. Reznikoff, M. Riley, M. Schaechter, and H. E. Umbarger (ed.), Escherichia coli and Salmonella:cellular and molecular biology, 2nd ed. ASM Press, Washington, D.C.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

