Group I self-splicing intron in the recA gene of Bacillus anthracis
- PMID: 12081963
- PMCID: PMC135159
- DOI: 10.1128/JB.184.14.3917-3922.2002
Group I self-splicing intron in the recA gene of Bacillus anthracis
Abstract
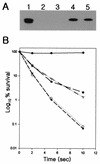
Self-splicing introns are rarely found in bacteria and bacteriophages. They are classified into group I and II according to their structural features and splicing mechanisms. While the group I introns are occasionally found in protein-coding regions of phage genomes and in several tRNA genes of cyanobacteria and proteobacteria, they had not been found in protein-coding regions of bacterial genomes. Here we report a group I intron in the recA gene of Bacillus anthracis which was initially found by DNA sequencing as an intervening sequence (IVS). By using reverse transcriptase PCR, the IVS was shown to be removable from the recA precursor mRNA for RecA that was being translated in E. coli. The splicing was visualized in vitro with labeled free GTP, indicating that it is a group I intron, which is also implied by its predicted secondary structure. The RecA protein of B. anthracis expressed in E. coli was functional in its ability to complement a recA defect. When recA-negative E. coli cells were irradiated with UV, the Bacillus RecA reduced the UV susceptibility of the recA mutant, regardless of the presence of intron.
Figures
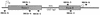
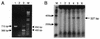

References
-
- Cech, T. R. 1990. Self-splicing of group I introns. Annu. Rev. Biochem. 59:543-568. - PubMed
-
- Davies, R. W., R. B. Waring, J. A. Ray, T. A. Brown, and C. Scazzocchio. 1982. Making ends meet: a model for RNA splicing in fungal mitochondria. Nature 300:719-724. - PubMed
-
- Davis, E. O., P. J. Jenner, P. C. Brooks, M. J. Colston, and S. G. Sedgwick. 1992. Protein splicing in the maturation of M. tuberculosis RecA protein: a mechanism for tolerating a novel class of intervening sequence. Cell 71:201-210. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

