Live-cell imaging reveals divergent intracellular dynamics of polyglutamine disease proteins and supports a sequestration model of pathogenesis
- PMID: 12084819
- PMCID: PMC123137
- DOI: 10.1073/pnas.152101299
Live-cell imaging reveals divergent intracellular dynamics of polyglutamine disease proteins and supports a sequestration model of pathogenesis
Abstract
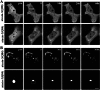
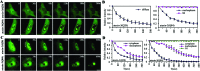
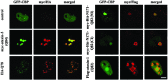
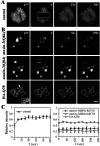
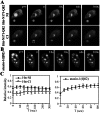
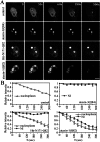
Protein misfolding and aggregation are central features of the polyglutamine neurodegenerative disorders, but the dynamic properties of expanded polyglutamine proteins are poorly understood. Here, we use fluorescence recovery after photobleaching (FRAP) and fluorescence loss in photobleaching (FLIP) with green fluorescent protein fusion proteins to study polyglutamine protein kinetics in living cells. Our results reveal markedly divergent mobility states for an expanded polyglutamine protein, ataxin-3, and establish that nuclear inclusions formed by this protein are aggregates. Additional studies of green fluorescent protein-tagged cAMP response element binding protein coexpressed with either of two mutant polyglutamine proteins, ataxin-3 and huntingtin, support a model of disease in which coaggregation of transcriptional components contributes to pathogenesis. Finally, studies of a third polyglutamine disease protein, ataxin-1, reveal unexpected heterogeneity in the dynamics of inclusions formed by different disease proteins, a finding which may help explain disease-specific elements of pathogenesis in these neurodegenerative disorders.
Figures
References
-
- Margolis R L, Ross C A. Trends Mol Med. 2001;7:479–482. - PubMed
-
- Cummings C J, Mancini M A, Antalffy B, DeFranco D B, Orr H T, Zoghbi H Y. Nat Genet. 1998;19:148–154. - PubMed
-
- Chai Y, Koppenhafer S L, Shoesmith S J, Perez M K, Paulson H L. Hum Mol Genet. 1999;8:673–682. - PubMed
-
- Stenoien D L, Cummings C J, Adams H P, Mancini M G, Patel K, DeMartino G N, Marcelli M, Weigel N L, Mancini M A. Hum Mol Genet. 1999;8:731–741. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

