Characterization of Salmonella serovars by PCR-single-strand conformation polymorphism analysis
- PMID: 12089246
- PMCID: PMC120578
- DOI: 10.1128/JCM.40.7.2346-2351.2002
Characterization of Salmonella serovars by PCR-single-strand conformation polymorphism analysis
Abstract
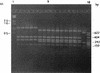
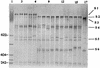
PCR-restriction fragment length polymorphism (PCR-RFLP) and PCR-single-strand conformation polymorphism (PCR-SSCP) analyses were carried out on the 1.6-kb groEL gene from 41 strains of 10 different Salmonella serovars. Three HaeIII RFLP profiles were recognized, but no discrimination between the serovars could be achieved by this technique. However, PCR-SSCP analysis of the groEL genes of various Salmonella serovars produced 14 SSCP profiles, indicating the potential of this technique to differentiate different Salmonella serovars (interserovar differentiation). Moreover, PCR-SSCP could differentiate strains within a subset of serovars (intraserovar discrimination), as three SSCP profiles were produced for the 11 Salmonella enterica serovar Enteritidis strains, and two SSCP profiles were generated for the 7 S. enterica serovar Infantis and five S. enterica serovar Newport strains. PCR-SSCP has the potential to complement classical typing methods such as serotyping and phage typing for the typing of Salmonella serovars due to its rapidity, simplicity, and typeability.
Figures
Similar articles
-
Rapid discrimination of Salmonella isolates by single-strand conformation polymorphism analysis.J Food Prot. 2008 Oct;71(10):1960-6. doi: 10.4315/0362-028x-71.10.1960. J Food Prot. 2008. PMID: 18939738
-
[Phylogenetic analysis and PCR-restriction fragment length polymophism identification of Salmonella based on grOEL gene sequence].Nan Fang Yi Ke Da Xue Xue Bao. 2009 Oct;29(10):2037-9, 2043. Nan Fang Yi Ke Da Xue Xue Bao. 2009. PMID: 19861260 Chinese.
-
Development of multiplex PCR for rapid identification of four Salmonella serovars most commonly isolated in Japan.Southeast Asian J Trop Med Public Health. 2014 May;45(3):654-61. Southeast Asian J Trop Med Public Health. 2014. PMID: 24974650
-
Salmonella serovars from foodborne and waterborne diseases in Korea, 1998-2007: total isolates decreasing versus rare serovars emerging.J Korean Med Sci. 2010 Dec;25(12):1693-9. doi: 10.3346/jkms.2010.25.12.1693. Epub 2010 Nov 24. J Korean Med Sci. 2010. PMID: 21165281 Free PMC article. Review.
-
Molecular methods for serovar determination of Salmonella.Crit Rev Microbiol. 2015;41(3):309-25. doi: 10.3109/1040841X.2013.837862. Epub 2013 Nov 14. Crit Rev Microbiol. 2015. PMID: 24228625 Review.
Cited by
-
One-step triplex high-resolution melting analysis for rapid identification and simultaneous subtyping of frequently isolated Salmonella serovars.Appl Environ Microbiol. 2012 May;78(9):3352-60. doi: 10.1128/AEM.07668-11. Epub 2012 Feb 17. Appl Environ Microbiol. 2012. PMID: 22344662 Free PMC article.
-
Differentiation of Erwinia amylovora and Erwinia pyrifoliae strains with single nucleotide polymorphisms and by synthesis of dihydrophenylalanine.Curr Microbiol. 2012 Jul;65(1):73-84. doi: 10.1007/s00284-012-0116-5. Epub 2012 Apr 27. Curr Microbiol. 2012. PMID: 22538467
-
Comparison of classical serotyping and PremiTest assay for routine identification of common Salmonella enterica serovars.J Clin Microbiol. 2008 Dec;46(12):4037-40. doi: 10.1128/JCM.01405-08. Epub 2008 Oct 8. J Clin Microbiol. 2008. PMID: 18842945 Free PMC article.
-
Microbial diversity of landslide soils assessed by RFLP and SSCP fingerprints.J Appl Genet. 2014 Aug;55(3):403-15. doi: 10.1007/s13353-014-0208-y. Epub 2014 Apr 11. J Appl Genet. 2014. PMID: 24722859
-
Aeromonas detection and characterization using genus-specific PCR and single-strand conformation polymorphism (SSCP).World J Microbiol Biotechnol. 2012 Oct;28(10):3007-13. doi: 10.1007/s11274-012-1111-5. Epub 2012 Jun 29. World J Microbiol Biotechnol. 2012. PMID: 22806741
References
-
- Aarts, H. J. M., L. A. J. T. Vanlith, and J. Keljer. 1998. High-resolution genotyping of Salmonella strains by AFLP fingerprinting. Lett. Appl. Microbiol. 26:131-135. - PubMed
-
- Cowan, S. T., and J. Steel. 1993. Cowan and Steel's manual for the identification of medical bacteria, 3rd ed. Cambridge University Press, Cambridge, United Kingdom.
-
- Ezquerra, E., A. Burnens, C. Jones, and J. Stanley. 1993. Genotypic typing and phylogenetic analysis of Salmonella paratyphi B and S. java with IS200. J. Gen. Microbiol. 139:2409-2414. - PubMed
-
- Gross, R. 1993., Signal transduction and virulence regulation in human and animal pathogens. FEMS Microbiol. Rev. 104:301-326. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

