Molecular peculiarities of the lytA gene isolated from clinical pneumococcal strains that are bile insoluble
- PMID: 12089276
- PMCID: PMC120542
- DOI: 10.1128/JCM.40.7.2545-2554.2002
Molecular peculiarities of the lytA gene isolated from clinical pneumococcal strains that are bile insoluble
Abstract
The autolytic LytA amidase from 12 bile (deoxycholate)-insoluble streptococcal isolates (formerly classified as atypical Streptococcus pneumoniae) showing different antibiotic resistance patterns was studied. These atypical strains, which autolyze at the end of the stationary phase of growth, contain highly divergent lytA alleles (pairwise evolutionary distances of about 20%) compared to the lytA alleles of typical pneumococci. The atypical LytA amidases exhibit a peculiar deletion of two amino acids responsible for cell wall anchoring in the carboxy-terminal domain and have a reduced specific activity. These enzymes were inhibited by 1% deoxycholate but were activated by 1% Triton X-100, a detergent that could be used as an alternative diagnostic test for this kind of strain. Preparation of functional chimeric enzymes, PCR mutagenesis, and gene replacements demonstrated that the characteristic bile insolubility of these atypical strains was due to their peculiar carboxy-terminal domain and that the 2-amino-acid deletion was responsible for the inhibitory effect of deoxycholate. However, the deletion alone did not affect the specific activity of LytA. A detailed characterization of the genes encoding the 16S rRNA and SodA together with multilocus sequence typing indicated that the strains studied here are not a single clone and, although they cannot be strictly classified as typical pneumococci, they represent a quite diverse pool of organisms closely related to S. pneumoniae. The clinical importance of these findings is underlined by the role of the lytA gene in shaping the course of pneumococcal diseases. This study can also contribute to solving diagnostic problems and to understanding the evolution and pathogenic potential of species of the Streptococcus mitis group.
Figures


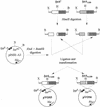




References
Publication types
MeSH terms
Substances
Associated data
- GDB/AJ419973
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

