Validation of DNA sequences using mass spectrometry coupled with nucleoside mass tagging
- PMID: 12097352
- PMCID: PMC186625
- DOI: 10.1101/gr.221402
Validation of DNA sequences using mass spectrometry coupled with nucleoside mass tagging
Abstract
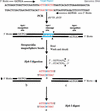
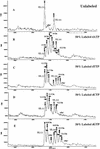
We present a mass spectrometry (MS)-based nucleoside-specific mass-tagging method to validate genomic DNA sequences containing ambiguities not resolved by gel electrophoresis. Selected types of (13)C/(15)N-labeled dNTPs are used in PCR amplification of target regions followed by matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF)-MS analysis. From the mass difference between the PCR products generated with unlabeled nucleosides and products containing (13)C/(15)N-labeled nucleosides, we determined the base composition of the genomic regions of interest. Two approaches were used to verify the target regions: The first approach used nucleosides partially enriched with stable isotopes to identify a single uncalled base in a gel electrophoresis-sequenced region. The second approach used mass tags with 100% heavy nucleosides to examine a GC-rich region of a polycytidine string with an unknown number of cytidines. By use of selected (13)C/(15)N-labeled dNTPs (dCTPs) in PCR amplification of the target region in tandem with MALDI-TOF-MS, we determined precisely that this string contains 11 cytidines. Both approaches show the ability of our MS-based mass-tagging strategy to solve critical questions of sequence identities that might be essential in determining the proper reading frames of the targeted regions.
Figures



References
-
- Blackstock WP, Weir MP. Proteomics: Quantitative and physical mapping of cellular proteins. Trends Biotechnol. 1999;17:121–127. - PubMed
-
- Chen X, Fei Z, Smith LM, Bradbury EM, Majidi V. Stable-isotope-assisted MALDI-TOF mass spectrometry for accurate determination of nucleotide compositions of PCR products. Anal Chem. 1999;71:3118–3128. - PubMed
-
- Fei Z, Smith LM. Analysis of single nucleotide polymorphisms by primer extension and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom. 2000;14:950–959. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
