The immunosuppressant rapamycin mimics a starvation-like signal distinct from amino acid and glucose deprivation
- PMID: 12101249
- PMCID: PMC133939
- DOI: 10.1128/MCB.22.15.5575-5584.2002
The immunosuppressant rapamycin mimics a starvation-like signal distinct from amino acid and glucose deprivation
Abstract
RAFT1/FRAP/mTOR is a key regulator of cell growth and division and the mammalian target of rapamycin, an immunosuppressive and anticancer drug. Rapamycin deprivation and nutrient deprivation have similar effects on the activity of S6 kinase 1 (S6K1) and 4E-BP1, two downstream effectors of RAFT1, but the relationship between nutrient- and rapamycin-sensitive pathways is unknown. Using transcriptional profiling, we show that, in human BJAB B-lymphoma cells and murine CTLL-2 T lymphocytes, rapamycin treatment affects the expression of many genes involved in nutrient and protein metabolism. The rapamycin-induced transcriptional profile is distinct from those induced by glucose, glutamine, or leucine deprivation but is most similar to that induced by amino acid deprivation. In particular, rapamycin treatment and amino acid deprivation up-regulate genes involved in nutrient catabolism and energy production and down-regulate genes participating in lipid and nucleotide synthesis and in protein synthesis, turnover, and folding. Surprisingly, however, rapamycin had effects opposite from those of amino acid starvation on the expression of a large group of genes involved in the synthesis, transport, and use of amino acids. Supported by measurements of nutrient use, the data suggest that RAFT1 is an energy and nutrient sensor and that rapamycin mimics a signal generated by the starvation of amino acids but that the signal is unlikely to be the absence of amino acids themselves. These observations underscore the importance of metabolism in controlling lymphocyte proliferation and offer a novel explanation for immunosuppression by rapamycin.
Figures



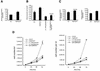
Similar articles
-
Assessment of cell-signaling pathways in the regulation of mammalian target of rapamycin (mTOR) by amino acids in rat adipocytes.J Cell Biochem. 2000 Sep 7;79(3):427-41. doi: 10.1002/1097-4644(20001201)79:3<427::aid-jcb80>3.0.co;2-0. J Cell Biochem. 2000. PMID: 10972980
-
RAFT1 phosphorylation of the translational regulators p70 S6 kinase and 4E-BP1.Proc Natl Acad Sci U S A. 1998 Feb 17;95(4):1432-7. doi: 10.1073/pnas.95.4.1432. Proc Natl Acad Sci U S A. 1998. PMID: 9465032 Free PMC article.
-
Regulation of targets of mTOR (mammalian target of rapamycin) signalling by intracellular amino acid availability.Biochem J. 2003 Jun 1;372(Pt 2):555-66. doi: 10.1042/BJ20021266. Biochem J. 2003. PMID: 12611592 Free PMC article.
-
Mechanism of action of the immunosuppressant rapamycin.Life Sci. 1996;58(5):373-95. doi: 10.1016/0024-3205(95)02233-3. Life Sci. 1996. PMID: 8594303 Review.
-
A novel pathway regulating the mammalian target of rapamycin (mTOR) signaling.Biochem Pharmacol. 2002 Oct 1;64(7):1071-7. doi: 10.1016/s0006-2952(02)01263-7. Biochem Pharmacol. 2002. PMID: 12234610 Review.
Cited by
-
Rapamycin enhances survival in a Drosophila model of mitochondrial disease.Oncotarget. 2016 Dec 6;7(49):80131-80139. doi: 10.18632/oncotarget.12560. Oncotarget. 2016. PMID: 27741510 Free PMC article.
-
Cytotoxicity of oleanane type triterpene from leaf extract of Pterospermum acerifolium (in vitro) and theoretical investigation of inhibitory signaling pathway.Chin Herb Med. 2020 Dec 1;13(1):124-130. doi: 10.1016/j.chmed.2020.09.004. eCollection 2021 Jan. Chin Herb Med. 2020. PMID: 36117756 Free PMC article.
-
Metabolic control by S6 kinases depends on dietary lipids.PLoS One. 2012;7(3):e32631. doi: 10.1371/journal.pone.0032631. Epub 2012 Mar 7. PLoS One. 2012. PMID: 22412899 Free PMC article.
-
Metabolism in T cell activation and differentiation.Curr Opin Immunol. 2010 Jun;22(3):314-20. doi: 10.1016/j.coi.2010.01.018. Epub 2010 Feb 26. Curr Opin Immunol. 2010. PMID: 20189791 Free PMC article. Review.
-
The integral role of mTOR in lipid metabolism.Cell Cycle. 2011 Mar 15;10(6):861-2. doi: 10.4161/cc.10.6.14930. Epub 2011 Mar 15. Cell Cycle. 2011. PMID: 21325894 Free PMC article. No abstract available.
References
-
- Barbosa-Tessmann, I. P., C. Chen, C. Zhong, F. Siu, S. M. Schuster, H. S. Nick, and M. S. Kilberg. 2000. Activation of the human asparagine synthetase gene by the amino acid response and the endoplasmic reticulum stress response pathways occurs by common genomic elements. J. Biol. Chem. 275:26976-26985. - PubMed
-
- Brown, E. J., M. W. Albers, T. B. Shin, K. Ichikawa, C. T. Keith, W. S. Lane, and S. L. Schreiber. 1994. A mammalian protein targeted by G1-arresting rapamycin-receptor complex. Nature 369:756-758. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
