Swarming-coupled expression of the Proteus mirabilis hpmBA haemolysin operon
- PMID: 12101306
- PMCID: PMC2528290
- DOI: 10.1099/00221287-148-7-2191
Swarming-coupled expression of the Proteus mirabilis hpmBA haemolysin operon
Abstract
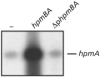
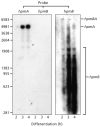
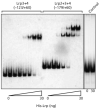
The HpmA haemolysin toxin of Proteus mirabilis is encoded by the hpmBA locus and its production is upregulated co-ordinately with the synthesis and assembly of flagella during differentiation into hyperflagellated swarm cells. Primer extension identified a sigma(70) promoter upstream of hpmB that was upregulated during swarming. Northern blotting indicated that this promoter region was also required for concomitant transcription of the immediately distal hpmA gene, and that the unstable hpmBA transcript generated a stable hpmA mRNA and an unstable hpmB mRNA. Transcriptional luxAB fusions to the DNA regions 5' of the hpmB and hpmA genes confirmed that hpmB sigma(70) promoter activity increased in swarm cells, and that there was no independent hpmA promoter. Increased transcription of the hpmBA operon in swarm cells was dependent upon a 125 bp sequence 5' of the sigma(70) promoter -35 hexamer. This sequence spans multiple putative binding sites for the leucine-responsive regulatory protein (Lrp), and band-shift assays with purified Lrp confirmed the presence of at least two such sites. The influence on hpmBA expression of the key swarming positive regulators FlhD(2)C(2) (encoded by the flagellar master operon), Lrp, and the membrane-located upregulator of the master operon, UmoB, was examined. Overexpression of each of these regulators moderately increased hpmBA transcription in wild-type P. mirabilis, and the hpmBA operon was not expressed in any of the flhDC, lrp or umoB mutants. Expression in the mutants was not recovered by cross-complementation, i.e. by overexpression of FlhD(2)C(2), Lrp or UmoB. Expression of the zapA protease virulence gene, which like hpmBA is also upregulated in swarm cells, did not require Lrp, but like flhDC it was upregulated by UmoB. The results indicate intersecting pathways of control linking virulence gene expression and swarm cell differentiation.
Figures


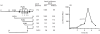

Similar articles
-
Novel genes that upregulate the Proteus mirabilis flhDC master operon controlling flagellar biogenesis and swarming.Mol Microbiol. 1998 Aug;29(3):741-51. doi: 10.1046/j.1365-2958.1998.00967.x. Mol Microbiol. 1998. PMID: 9723914
-
A nonswarming mutant of Proteus mirabilis lacks the Lrp global transcriptional regulator.J Bacteriol. 1997 Aug;179(15):4741-6. doi: 10.1128/jb.179.15.4741-4746.1997. J Bacteriol. 1997. PMID: 9244260 Free PMC article.
-
Functions of the subunits in the FlhD(2)C(2) transcriptional master regulator of bacterial flagellum biogenesis and swarming.J Mol Biol. 2000 Nov 3;303(4):467-78. doi: 10.1006/jmbi.2000.4149. J Mol Biol. 2000. PMID: 11054284
-
Requirement for FlhA in flagella assembly and swarm-cell differentiation by Proteus mirabilis.Mol Microbiol. 1995 Feb;15(4):761-9. doi: 10.1111/j.1365-2958.1995.tb02383.x. Mol Microbiol. 1995. PMID: 7783646
-
Regulation of gene expression during swarmer cell differentiation in Proteus mirabilis.FEMS Microbiol Rev. 2010 Sep;34(5):753-63. doi: 10.1111/j.1574-6976.2010.00229.x. Epub 2010 Apr 19. FEMS Microbiol Rev. 2010. PMID: 20497230 Review.
Cited by
-
Proteus mirabilis Biofilm: Development and Therapeutic Strategies.Front Cell Infect Microbiol. 2020 Aug 14;10:414. doi: 10.3389/fcimb.2020.00414. eCollection 2020. Front Cell Infect Microbiol. 2020. PMID: 32923408 Free PMC article. Review.
-
Vaccination with proteus toxic agglutinin, a hemolysin-independent cytotoxin in vivo, protects against Proteus mirabilis urinary tract infection.Infect Immun. 2009 Feb;77(2):632-41. doi: 10.1128/IAI.01050-08. Epub 2008 Nov 24. Infect Immun. 2009. PMID: 19029299 Free PMC article.
-
Molecular detection of HpmA and HlyA hemolysin of uropathogenic Proteus mirabilis.Curr Microbiol. 2013 Dec;67(6):703-7. doi: 10.1007/s00284-013-0423-5. Epub 2013 Jul 25. Curr Microbiol. 2013. PMID: 23884594
-
The acetate switch.Microbiol Mol Biol Rev. 2005 Mar;69(1):12-50. doi: 10.1128/MMBR.69.1.12-50.2005. Microbiol Mol Biol Rev. 2005. PMID: 15755952 Free PMC article. Review.
-
Complicated catheter-associated urinary tract infections due to Escherichia coli and Proteus mirabilis.Clin Microbiol Rev. 2008 Jan;21(1):26-59. doi: 10.1128/CMR.00019-07. Clin Microbiol Rev. 2008. PMID: 18202436 Free PMC article. Review.
References
-
- Akiyama M, Horiuchi T, Sekiguchi M. Molecular cloning and nucleotide sequence of the mutT mutator of Escherichia coli that causes A:T to C:G transversion. Mol Gen Genet. 1987;206:9–16. - PubMed
-
- Allison C, Hughes C. Closely linked genetic loci required for swarm cell differentiation and multicellular migration by Proteus mirabilis. Mol Microbiol. 1991;5:1975–1982. - PubMed
-
- Allison C, Lai H-C, Hughes C. Co-ordinate expression of virulence genes during swarm-cell differentiation and population migration of Proteus mirabilis. Mol Microbiol. 1992;6:1583–1591. - PubMed
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman D. Basic logical alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl L. Current Protocols in Molecular Biology. New York: Wiley; 1988.
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

