Transcription factor-mediated lineage switching reveals plasticity in primary committed progenitor cells
- PMID: 12110589
- PMCID: PMC126114
- DOI: 10.1093/emboj/cdf368
Transcription factor-mediated lineage switching reveals plasticity in primary committed progenitor cells
Abstract
The developmental plasticity of transplanted adult stem cells challenges the notion that tissue-restricted stem cells have stringently limited lineage potential and prompts a re-evaluation of the stability of lineage commitment. Transformed cell systems are inappropriate for such studies, since transformation potentially dysregulates the processes governing lineage commitment. We have therefore assessed the stability of normal lineage commitment in primary adult haematopoietic cells. For these studies we have used prospectively isolated primary bipotent progenitors, which normally display only neutrophil and monocyte differentiation in vitro. In response to ectopic transcription factor expression, these neutrophil/monocyte progenitors were reprogrammed to take on erythroid, eosinophil and basophil-like cell fates, with the resultant colonies resembling the mixed lineage colonies normally generated by multipotential progenitors. Clone-marking and daughter cell experiments identified lineage switching rather than differential cell selection as the mechanism of altered lineage output. These results demonstrate that the cell type-specific programming of apparently committed primary progenitors is not irrevocably fixed, but may be radically re-specified in response to a single transcriptional regulator.
Figures
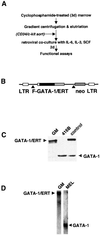
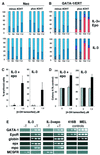





References
-
- Akashi K., Traver,D., Miyamoto,T. and Weissman,I.L. (2000) A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature, 404, 193–197. - PubMed
-
- Anderson D.J., Gage,F.H. and Weissman,I.L. (2001) Can stem cells cross lineage boundaries? Nat. Med., 7, 393–395. - PubMed
-
- Choi J., Costa,M.L., Mermelstein,C.S., Chagas,C., Holtzer,S. and Holtzer,H. (1990) MyoD converts primary dermal fibroblasts, chondroblasts, smooth muscle, and retinal pigmented epithelial cells into striated mononucleated myoblasts and multinucleated myotubes. Proc. Natl Acad. Sci. USA, 87, 7988–7992. - PMC - PubMed
-
- Cook N., Dexter,T.M., Lord,B.I., Cragoe,E.J.,Jr and Whetton,A.D. (1989) Identification of a common signal associated with cellular proliferation stimulated by four haemopoietic growth factors in a highly enriched population of granulocyte/macrophage colony-forming cells. EMBO J., 8, 2967–2974. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

