Replication fork collapse at replication terminator sequences
- PMID: 12110601
- PMCID: PMC126115
- DOI: 10.1093/emboj/cdf369
Replication fork collapse at replication terminator sequences
Abstract
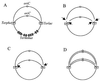
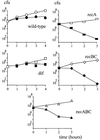
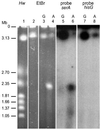
Replication fork arrest is a source of genome re arrangements, and the recombinogenic properties of blocked forks are likely to depend on the cause of blockage. Here we study the fate of replication forks blocked at natural replication arrest sites. For this purpose, Escherichia coli replication terminator sequences Ter were placed at ectopic positions on the bacterial chromosome. The resulting strain requires recombinational repair for viability, but replication forks blocked at Ter are not broken. Linear DNA molecules are formed upon arrival of a second round of replication forks that copy the DNA strands of the first blocked forks to the end. A model that accounts for the requirement for homologous recombination for viability in spite of the lack of chromosome breakage is proposed. This work shows that natural and accidental replication arrests sites are processed differently.
Figures
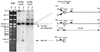
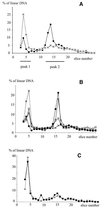
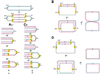
Similar articles
-
The Escherichia coli UvrD helicase is essential for Tus removal during recombination-dependent replication restart from Ter sites.Mol Microbiol. 2006 Oct;62(2):382-96. doi: 10.1111/j.1365-2958.2006.05382.x. Mol Microbiol. 2006. PMID: 17020578
-
Mechanisms of polar arrest of a replication fork.Mol Microbiol. 2009 Apr;72(2):279-85. doi: 10.1111/j.1365-2958.2009.06656.x. Epub 2009 Mar 4. Mol Microbiol. 2009. PMID: 19298368 Review.
-
Requirement for RecFOR-mediated recombination in priA mutant.Mol Microbiol. 2004 Apr;52(2):551-62. doi: 10.1111/j.1365-2958.2004.03997.x. Mol Microbiol. 2004. PMID: 15066040
-
Replication termination in Escherichia coli: structure and antihelicase activity of the Tus-Ter complex.Microbiol Mol Biol Rev. 2005 Sep;69(3):501-26. doi: 10.1128/MMBR.69.3.501-526.2005. Microbiol Mol Biol Rev. 2005. PMID: 16148308 Free PMC article. Review.
-
Instability of inhibited replication forks in E. coli.Bioessays. 1995 Aug;17(8):733-41. doi: 10.1002/bies.950170810. Bioessays. 1995. PMID: 7661854 Review.
Cited by
-
Shaping the landscape of the Escherichia coli chromosome: replication-transcription encounters in cells with an ectopic replication origin.Nucleic Acids Res. 2015 Sep 18;43(16):7865-77. doi: 10.1093/nar/gkv704. Epub 2015 Jul 8. Nucleic Acids Res. 2015. PMID: 26160884 Free PMC article.
-
Tracking of controlled Escherichia coli replication fork stalling and restart at repressor-bound DNA in vivo.EMBO J. 2006 Jun 7;25(11):2596-604. doi: 10.1038/sj.emboj.7601155. Epub 2006 May 25. EMBO J. 2006. PMID: 16724111 Free PMC article.
-
Interplay between chromosomal architecture and termination of DNA replication in bacteria.Front Microbiol. 2023 Jun 26;14:1180848. doi: 10.3389/fmicb.2023.1180848. eCollection 2023. Front Microbiol. 2023. PMID: 37434703 Free PMC article. Review.
-
Growth rate toxicity phenotypes and homeostatic supercoil control differentiate Escherichia coli from Salmonella enterica serovar Typhimurium.J Bacteriol. 2007 Aug;189(16):5839-49. doi: 10.1128/JB.00083-07. Epub 2007 Mar 30. J Bacteriol. 2007. PMID: 17400739 Free PMC article.
-
Direct restart of a replication fork stalled by a head-on RNA polymerase.J Vis Exp. 2010 Apr 29;(38):1919. doi: 10.3791/1919. J Vis Exp. 2010. PMID: 20436399 Free PMC article.
References
-
- Aguilera A., Chavez,S. and Malagon,F. (2000) Mitotic recombination in yeast: elements controlling its incidence. Yeast, 16, 731–754. - PubMed
-
- Bierne H., Vilette,D., Ehrlich,S.D. and Michel,B. (1997b) Isolation of a dnaE mutation which enhances RecA-independent homologous recombination in the Escherichia coli chromosome. Mol. Microbiol., 24, 1225–1234. - PubMed
-
- Bussiere D.E. and Bastia,D. (1999) Termination of DNA replication of bacterial and plasmid chromosomes. Mol. Microbiol., 31, 1611–1618. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

