Regulation of squalene synthase, a key enzyme of sterol biosynthesis, in tobacco
- PMID: 12114564
- PMCID: PMC166504
- DOI: 10.1104/pp.001438
Regulation of squalene synthase, a key enzyme of sterol biosynthesis, in tobacco
Abstract
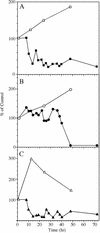
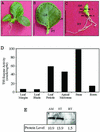
Squalene synthase (SS) represents a putative branch point in the isoprenoid biosynthetic pathway capable of diverting carbon flow specifically to the biosynthesis of sterols and, hence, is considered a potential regulatory point for sterol metabolism. For example, when plant cells grown in suspension culture are challenged with fungal elicitors, suppression of sterol biosynthesis has been correlated with a reduction in SS enzyme activity. The current study sought to correlate changes in SS enzyme activity with changes in the level of the corresponding protein and mRNA. Using an SS-specific antibody, the initial suppression of SS enzyme activity in elicitor-challenged cells was not reflected by changes in the absolute level of the corresponding polypeptide, implicating a post-translational control mechanism for this enzyme activity. In comparison, the absolute level of the SS mRNA did decrease approximately 5-fold in the elicitor-treated cells, which is suggestive of decreased transcription of the SS gene. Study of SS in intact plants was also initiated by measuring the level of SS enzyme activity, the level of the corresponding protein, and the expression of SS gene promoter-reporter gene constructs in transgenic plants. SS enzyme activity, polypeptide level, and gene expression were all localized predominately to the shoot apical meristem, with much lower levels observed in leaves and roots. These later results suggest that sterol biosynthesis is localized to the apical meristems and that apical meristems may be a source of sterols for other plant tissues.
Figures



References
-
- Adler JH, Grebenok RJ. Biosynthesis and distribution of insect-molting hormones in plants: a review. Lipids. 1995;30:257–262. - PubMed
-
- Bhatt PN, Bhatt DP. Regulation of sterol biosynthesis in Solanum species. J Exp Bot. 1984;35:890–896.
-
- Brindle PA, Kuhn PJ, Threlfall DR. Biosynthesis and metabolism of sesquiterpenoid phytoalexins and triterpenoids in potato cell-suspension cultures. Phytochemistry. 1988;27:133–150.
-
- Brown MS, Goldstein JL. The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell. 1997;89:331–340. - PubMed
-
- Caelles C, Ferrer A, Balcells L, Hegardt FG, Boronat A. Isolation and structural characterization of a cDNA-encoding Arabidopsis thaliana 3-hydroxy-3-methylglutaryl coenzyme-A reductase. Plant Mol Biol. 1989;13:627–638. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

