Microarray-based identification of htrA, a Streptococcus pneumoniae gene that is regulated by the CiaRH two-component system and contributes to nasopharyngeal colonization
- PMID: 12117912
- PMCID: PMC128155
- DOI: 10.1128/IAI.70.8.4059-4067.2002
Microarray-based identification of htrA, a Streptococcus pneumoniae gene that is regulated by the CiaRH two-component system and contributes to nasopharyngeal colonization
Abstract
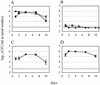
Nasopharyngeal carriage is the reservoir from which most disease with Streptococcus pneumoniae arises. Survival as a commensal in this environment is likely to require a set of adaptations distinct from those needed to cause disease, some of which may be mediated by two-component signal transduction systems (TCSTS). We examined the contributions of nine pneumococcal TCSTS to the process of nasopharyngeal colonization by using an infant rat model. Whereas deletions in all but one of these systems have been associated previously with a high degree of attenuation in a murine model of pneumonia, only the CiaRH system was necessary for efficient carriage. Transcriptional analysis by using microarray hybridization identified a locus consisting of two adjacent genes, htrA and spoJ, that was specifically and strongly downregulated in a DeltaciaRH-null mutant. A S. pneumoniae strain lacking the htrA gene encoding a putative serine protease, but not one lacking spoJ, showed decreased fitness in a competitive model of colonization, a finding consistent with this gene mediating a portion of the carriage deficit observed with the DeltaciaRH strain.
Figures

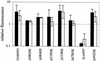

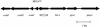
Similar articles
-
Bacteriocin activity of Streptococcus pneumoniae is controlled by the serine protease HtrA via posttranscriptional regulation.J Bacteriol. 2009 Mar;191(5):1509-18. doi: 10.1128/JB.01213-08. Epub 2008 Dec 19. J Bacteriol. 2009. PMID: 19103930 Free PMC article.
-
Pneumococcal HtrA protease mediates inhibition of competence by the CiaRH two-component signaling system.J Bacteriol. 2005 Jun;187(12):3969-79. doi: 10.1128/JB.187.12.3969-3979.2005. J Bacteriol. 2005. PMID: 15937159 Free PMC article.
-
Control of virulence by the two-component system CiaR/H is mediated via HtrA, a major virulence factor of Streptococcus pneumoniae.J Bacteriol. 2004 Aug;186(16):5258-66. doi: 10.1128/JB.186.16.5258-5266.2004. J Bacteriol. 2004. PMID: 15292127 Free PMC article.
-
Influence of bacterial interactions on pneumococcal colonization of the nasopharynx.Trends Microbiol. 2013 Mar;21(3):129-35. doi: 10.1016/j.tim.2012.11.005. Epub 2012 Dec 25. Trends Microbiol. 2013. PMID: 23273566 Free PMC article. Review.
-
The HtrA family of serine proteases.Mol Microbiol. 1997 Oct;26(2):209-21. doi: 10.1046/j.1365-2958.1997.5601928.x. Mol Microbiol. 1997. PMID: 9383148 Review.
Cited by
-
Bacteriocin activity of Streptococcus pneumoniae is controlled by the serine protease HtrA via posttranscriptional regulation.J Bacteriol. 2009 Mar;191(5):1509-18. doi: 10.1128/JB.01213-08. Epub 2008 Dec 19. J Bacteriol. 2009. PMID: 19103930 Free PMC article.
-
Streptococcus pneumoniae Senses a Human-like Sialic Acid Profile via the Response Regulator CiaR.Cell Host Microbe. 2016 Sep 14;20(3):307-317. doi: 10.1016/j.chom.2016.07.019. Epub 2016 Sep 1. Cell Host Microbe. 2016. PMID: 27593514 Free PMC article.
-
Highly Variable Streptococcus oralis Strains Are Common among Viridans Streptococci Isolated from Primates.mSphere. 2016 Mar 9;1(2):e00041-15. doi: 10.1128/mSphere.00041-15. eCollection 2016 Mar-Apr. mSphere. 2016. PMID: 27303717 Free PMC article.
-
Variation in the presence of neuraminidase genes among Streptococcus pneumoniae isolates with identical sequence types.Infect Immun. 2006 Jun;74(6):3360-5. doi: 10.1128/IAI.01442-05. Infect Immun. 2006. PMID: 16714565 Free PMC article.
-
Streptococcus pneumoniae nasopharyngeal colonization induces type I interferons and interferon-induced gene expression.BMC Genomics. 2009 Aug 27;10:404. doi: 10.1186/1471-2164-10-404. BMC Genomics. 2009. PMID: 19712482 Free PMC article.
References
-
- Bartilson, M., A. Marra, J. Christine, J. S. Asundi, W. P. Schneider, and A. E. Hromockyj. 2001. Differential fluorescence induction reveals Streptococcus pneumoniae loci regulated by competence stimulatory peptide. Mol. Microbiol. 39:126-135. - PubMed
-
- Chatfield, S. N., K. Strahan, D. Pickard, I. G. Charles, C. E. Hormaeche, and G. Dougan. 1992. Evaluation of Salmonella typhimurium strains harbouring defined mutations in htrA and aroA in the murine salmonellosis model. Microb. Pathog. 12:145-151. - PubMed
-
- Cheng, Q., E. A. Campbell, A. M. Naughton, S. Johnson, and H. R. Masure. 1997. The com locus controls genetic transformation in Streptococcus pneumoniae. Mol. Microbiol. 23:683-692. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

