Yersiniabactin production requires the thioesterase domain of HMWP2 and YbtD, a putative phosphopantetheinylate transferase
- PMID: 12117929
- PMCID: PMC128149
- DOI: 10.1128/IAI.70.8.4204-4214.2002
Yersiniabactin production requires the thioesterase domain of HMWP2 and YbtD, a putative phosphopantetheinylate transferase
Abstract
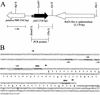
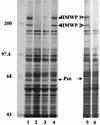
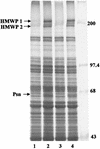
One requirement for the pathogenesis of Yersinia pestis, the causative agent of bubonic plague, is the yersiniabactin (Ybt) siderophore-dependent iron transport system that is encoded within a high-pathogenicity island (HPI) within the pgm locus of the Y. pestis chromosome. Nine gene products within the HPI have demonstrated functions in the nonribosomal peptide synthesis (NRPS)/polyketide (PK) synthesis or transport of Ybt. NRPS/PK synthetase or synthase enzymes are generally activated by phosphopantetheinylation. However, no products with similarities to known phosphopantetheinyl (P-pant) transferases were found within the pgm locus. We have identified a gene, ybtD, encoded outside the HPI and pgm locus, that is necessary for function of the Ybt system and has similarities to other P-pant transferases such as EntD of Escherichia coli. A deletion within ybtD yielded a strain (KIM6-2085+) defective in siderophore production. This strain was unable to grow on iron-deficient media at 37 degrees C but could be cross-fed by culture supernatants from Ybt-producing strains of Y. pestis. The promoter region of ybtD was fused to lacZ; beta-galactosidase expression from this reporter was not regulated by the iron status of the bacterial cells or by YbtA, a positive regulator of other genes of the ybt system. The ybtD mutant failed to express indicator Ybt proteins (high-molecular-weight protein 1 [HMWP1], HMWP2, and Psn), a pattern similar to those seen with several other ybt biosynthetic mutants. In contrast, cells containing a single amino acid substitution (S2908A) in the terminal thioesterase domain of HMWP2 failed to exhibit any ybt regulatory defects but did not elaborate extracellular Ybt under iron-deficient conditions.
Figures
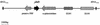


References
-
- Armstrong, S. K., G. S. Pettis, L. J. Forrester, and M. A. McIntosh. 1989. The Escherichia coli enterobactin biosynthesis gene, entD: nucleotide sequence and membrane localization of its protein product. Mol. Microbiol. 3:757-766. - PubMed
-
- Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl (ed.). 1987. Current protocols in molecular biology. John Wiley & Sons, New York, N.Y.
-
- Bearden, S. W., and R. D. Perry. 1999. The Yfe system of Yersinia pestis transports iron and manganese and is required for full virulence of plague. Mol. Microbiol. 32:403-414. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

