Role of polyphosphate kinase in biofilm formation by Porphyromonas gingivalis
- PMID: 12117989
- PMCID: PMC128176
- DOI: 10.1128/IAI.70.8.4708-4715.2002
Role of polyphosphate kinase in biofilm formation by Porphyromonas gingivalis
Abstract
In order to assess the role of polyphosphate kinase (PPK) in the physiology of Porphyromonas gingivalis, a ppk gene mutant, CW120, was constructed and characterized. P. gingivalis was demonstrated to synthesize short-chain polyphosphate (polyP) but not long-chain polyP. CW120 failed to survive in the stationary phase as well as the parental cell did, and it was attenuated in biofilm formation on polyvinylchloride and glass surfaces. Furthermore, the complementation by insertion of an intact copy of the ppk gene into the mutant CW120 restored its biofilm formation and stationary-phase survival. These results suggest that PPK may be important for incorporation of these organisms into subgingival plaque in the human oral cavity.
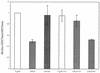
Figures




Similar articles
-
Biofilm formation by the periodontopathic bacteria Treponema denticola and Porphyromonas gingivalis.J Periodontol. 2005 Nov;76(11 Suppl):2047-51. doi: 10.1902/jop.2005.76.11-S.2047. J Periodontol. 2005. PMID: 16277575
-
Genetically altered levels of inorganic polyphosphate in Escherichia coli.J Biol Chem. 1994 Mar 4;269(9):6290-5. J Biol Chem. 1994. PMID: 8119977
-
Polyphosphate kinase of Lysinibacillus sphaericus and its effects on accumulation of polyphosphate and bacterial growth.Microbiol Res. 2015 Mar;172:41-7. doi: 10.1016/j.micres.2014.12.002. Epub 2014 Dec 12. Microbiol Res. 2015. PMID: 25541179
-
Inorganic polyphosphate and polyphosphate kinase: their novel biological functions and applications.Biochemistry (Mosc). 2000 Mar;65(3):315-23. Biochemistry (Mosc). 2000. PMID: 10739474 Review.
-
Dual lifestyle of Porphyromonas gingivalis in biofilm and gingival cells.Microb Pathog. 2016 May;94:42-7. doi: 10.1016/j.micpath.2015.10.003. Epub 2015 Oct 9. Microb Pathog. 2016. PMID: 26456558 Review.
Cited by
-
Polyphosphate deficiency in Mycobacterium tuberculosis is associated with enhanced drug susceptibility and impaired growth in guinea pigs.J Bacteriol. 2013 Jun;195(12):2839-51. doi: 10.1128/JB.00038-13. Epub 2013 Apr 12. J Bacteriol. 2013. PMID: 23585537 Free PMC article.
-
Polyphosphate kinase 1 is a pathogenesis determinant in Campylobacter jejuni.J Bacteriol. 2007 Nov;189(22):8099-108. doi: 10.1128/JB.01037-07. Epub 2007 Sep 7. J Bacteriol. 2007. PMID: 17827292 Free PMC article.
-
Polyphosphate Plays a Significant Role in the Maturation of Spores in Myxococcus xanthus.Curr Microbiol. 2024 Jul 1;81(8):248. doi: 10.1007/s00284-024-03778-7. Curr Microbiol. 2024. PMID: 38951187
-
Colistin-resistant Pseudomonas aeruginosa clinical strains with defective biofilm formation.GMS Hyg Infect Control. 2019 Oct 10;14:Doc12. doi: 10.3205/dgkh000328. eCollection 2019. GMS Hyg Infect Control. 2019. PMID: 31728266 Free PMC article.
-
Role of Polyphosphate in Amyloidogenic Processes.Cold Spring Harb Perspect Biol. 2019 May 1;11(5):a034041. doi: 10.1101/cshperspect.a034041. Cold Spring Harb Perspect Biol. 2019. PMID: 30617049 Free PMC article. Review.
References
-
- Ahn, K., and A. Kornberg. 1990. Polyphosphate kinase from Escherichia coli. Purification and demonstration of a phosphoenzyme intermediate. J. Biol. Chem. 265:11734-11739. - PubMed
-
- Akiyama, M., E. Crooke, and A. Kornberg. 1993. An exopolyphosphatase of Escherichia coli. The enzyme and its ppx gene in a polyphosphate operon. J. Biol. Chem. 268:633-639. - PubMed
-
- Akiyama, M., E. Crooke, and A. Kornberg. 1992. The polyphosphate kinase gene of Escherichia coli. Isolation and sequence of the ppk gene and membrane location of the protein. J. Biol. Chem. 267:22556-22561. - PubMed
-
- Booth, J. W., and G. Guidotti. 1995. An alleged yeast polyphosphate kinase is actually diadenosine-5′,5″-P1, P4-tetraphosphate alpha,beta-phosphorylase. J. Biol. Chem. 270:19377-19382. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

