Favorable and unfavorable HLA class I alleles and haplotypes in Zambians predominantly infected with clade C human immunodeficiency virus type 1
- PMID: 12134033
- PMCID: PMC155130
- DOI: 10.1128/jvi.76.16.8276-8284.2002
Favorable and unfavorable HLA class I alleles and haplotypes in Zambians predominantly infected with clade C human immunodeficiency virus type 1
Abstract
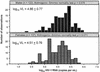
The setpoint of viral RNA concentration (viral load [VL]) during chronic human immunodeficiency virus type 1 (HIV-1) infection reflects a virus-host equilibration closely related to CD8(+) cytotoxic T-lymphocyte (CTL) responses, which rely heavily on antigen presentation by the human major histocompatibility complex (MHC) (i.e., HLA) class I molecules. Differences in HIV-1 VL among 259 mostly clade C virus-infected individuals (137 females and 122 males) in the Zambia-UAB HIV Research Project (ZUHRP) were associated with several HLA class I alleles and haplotypes. In particular, general linear model analyses revealed lower log(10) VL among those with HLA allele B*57 (P = 0.002 [without correction]) previously implicated in favorable response and in those with HLA B*39 and A*30-Cw*03 (P = 0.002 to 0.016); the same analyses also demonstrated higher log(10) VL among individuals with A*02-Cw*16, A*23-B*14, and A*23-Cw*07 (P = 0.010 to 0.033). These HLA effects remained strong (P = 0.0002 to 0.075) after adjustment for age, gender, and duration of infection and persisted across three orders of VL categories (P = 0.001 to 0.084). In contrast, neither B*35 (n = 15) nor B*53 (n = 53) showed a clear disadvantage such as that reported elsewhere for these closely related alleles. Other HLA associations with unusually high (A*68, B*41, B*45, and Cw*16) or low (B*13, Cw*12, and Cw*18) VL were either unstable or reflected their tight linkage respecting disequilibria with other class I variants. The three consistently favorable HLA class I variants retained in multivariable models and in alternative analyses were present in 30.9% of subjects with the lowest (<10,000 copies per ml) and 3.1% of those with the highest (>100,000) VL. Clear differential distribution of HLA profiles according to level of viremia suggests important host genetic contribution to the pattern of immune control and escape during HIV-1 infection.
Figures
References
-
- Balla-Jhagjhoorsingh, S. S., G. Koopman, P. Mooij, T. G. M. Haaksman, V. J. P. Teeuwsen, R. E. Bontrop, and J. L. Neeney. 1999. Conserved CTL epitope shared between HIV-infected human long-term survivors and chimpanzees. J. Immunol. 162:2308-2314. - PubMed
-
- Bienzle, D., K. S. MacDonald, F. M. Smaill, C. Kovacs, M. Baqi, B. Courssaris, M. A. Luscher, S. L. Walmsley, and K. L. Rosenthal. 2000. Factors contributing to the lack of human immunodeficiency virus type 1 (HIV-1) transmission in HIV-1-discordant partners. J. Infect. Dis. 182:123-132. - PubMed
-
- Bratt, G., A. Karlsson, A. C. Leandersson, J. Albert, B. Wahren, and E. Sandstrom. 1998. Treatment history and baseline viral load, but not viral tropism or CCR-5 genotype, influence prolonged antiviral efficacy of highly active antiretroviral treatment. AIDS 12:2193-2202. - PubMed
-
- Cao, K., J. Hollenbach, X. Shi, W. Shi, M. Chopek, and M. A. Fernandez-Vina. 2001. Analysis of the frequencies of HLA-A, B, and C alleles and haplotypes in the five major ethnic groups of the United States reveals high levels of diversity in these loci and contrasting distribution patterns in these populations. Hum. Immunol. 62:1009-1030. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials