Mammalian reovirus nonstructural protein microNS forms large inclusions and colocalizes with reovirus microtubule-associated protein micro2 in transfected cells
- PMID: 12134034
- PMCID: PMC155143
- DOI: 10.1128/jvi.76.16.8285-8297.2002
Mammalian reovirus nonstructural protein microNS forms large inclusions and colocalizes with reovirus microtubule-associated protein micro2 in transfected cells
Abstract
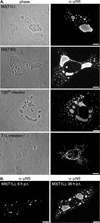
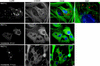
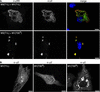
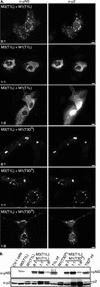
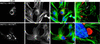
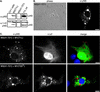
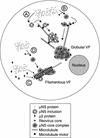
Cells infected with mammalian orthoreoviruses contain large cytoplasmic phase-dense inclusions believed to be the sites of viral replication and assembly, but the morphogenesis, structure, and specific functions of these "viral factories" are poorly understood. Using immunofluorescence microscopy, we found that reovirus nonstructural protein microNS expressed in transfected cells forms inclusions that resemble the globular viral factories formed in cells infected with reovirus strain type 3 Dearing from our laboratory (T3D(N)). In the transfected cells, the formation of microNS large globular perinuclear inclusions was dependent on the microtubule network, as demonstrated by the appearance of many smaller microNS globular inclusions dispersed throughout the cytoplasm after treatment with the microtubule-depolymerizing drug nocodazole. Coexpression of microNS and reovirus protein micro2 from a different strain, type 1 Lang (T1L), which forms filamentous viral factories, altered the distributions of both proteins. In cotransfected cells, the two proteins colocalized in thick filamentous structures. After nocodazole treatment, many small dispersed globular inclusions containing microNS and micro2 were seen, demonstrating that the microtubule network is required for the formation of the filamentous structures. When coexpressed, the micro2 protein from T3D(N) also colocalized with microNS, but in globular inclusions rather than filamentous structures. The morphology difference between the globular inclusions containing microNS and micro2 protein from T3D(N) and the filamentous structures containing microNS and micro2 protein from T1L in cotransfected cells mimicked the morphology difference between globular and filamentous factories in reovirus-infected cells, which is determined by the micro2-encoding M1 genome segment. We found that the first 40 amino acids of microNS are required for colocalization with micro2 but not for inclusion formation. Similarly, a fusion of microNS amino acids 1 to 41 to green fluorescent protein was sufficient for colocalization with the micro2 protein from T1L but not for inclusion formation. These observations suggest a functional difference between microNS and microNSC, a smaller form of the protein that is present in infected cells and that is missing amino acids from the amino terminus of microNS. The capacity of microNS to form inclusions and to colocalize with micro2 in transfected cells suggests a key role for microNS in forming viral factories in reovirus-infected cells.
Figures


Similar articles
-
Reovirus core protein mu2 determines the filamentous morphology of viral inclusion bodies by interacting with and stabilizing microtubules.J Virol. 2002 May;76(9):4483-96. doi: 10.1128/jvi.76.9.4483-4496.2002. J Virol. 2002. PMID: 11932414 Free PMC article.
-
Reovirus mu2 protein determines strain-specific differences in the rate of viral inclusion formation in L929 cells.Virology. 2000 Jun 20;272(1):16-26. doi: 10.1006/viro.2000.0362. Virology. 2000. PMID: 10873745
-
Avian reovirus nonstructural protein microNS forms viroplasm-like inclusions and recruits protein sigmaNS to these structures.Virology. 2004 Feb 5;319(1):94-106. doi: 10.1016/j.virol.2003.10.034. Virology. 2004. PMID: 14967491
-
Function, Architecture, and Biogenesis of Reovirus Replication Neoorganelles.Viruses. 2019 Mar 21;11(3):288. doi: 10.3390/v11030288. Viruses. 2019. PMID: 30901959 Free PMC article. Review.
-
Early steps in avian reovirus morphogenesis.Curr Top Microbiol Immunol. 2006;309:67-85. doi: 10.1007/3-540-30773-7_3. Curr Top Microbiol Immunol. 2006. PMID: 16909897 Review.
Cited by
-
Aquareovirus NS80 recruits viral proteins to its inclusions, and its C-terminal domain is the primary driving force for viral inclusion formation.PLoS One. 2013;8(2):e55334. doi: 10.1371/journal.pone.0055334. Epub 2013 Feb 12. PLoS One. 2013. PMID: 23424630 Free PMC article.
-
Characterization of grass carp reovirus minor core protein VP4.Virol J. 2012 Jul 5;9:89. doi: 10.1186/1743-422X-9-89. Virol J. 2012. PMID: 22559058 Free PMC article.
-
A versatile molecular tagging method for targeting proteins to avian reovirus muNS inclusions. Use in protein immobilization and purification.PLoS One. 2010 Nov 12;5(11):e13961. doi: 10.1371/journal.pone.0013961. PLoS One. 2010. PMID: 21103063 Free PMC article.
-
Strain-specific differences in reovirus infection of murine macrophages segregate with polymorphisms in viral outer-capsid protein σ3.J Virol. 2024 Nov 19;98(11):e0114724. doi: 10.1128/jvi.01147-24. Epub 2024 Oct 21. J Virol. 2024. PMID: 39431846 Free PMC article.
-
Rotavirus viroplasm fusion and perinuclear localization are dynamic processes requiring stabilized microtubules.PLoS One. 2012;7(10):e47947. doi: 10.1371/journal.pone.0047947. Epub 2012 Oct 23. PLoS One. 2012. PMID: 23110139 Free PMC article.
References
-
- Antczak, J. B., and W. K. Joklik. 1992. Reovirus genome segment assortment into progeny genomes studied by the use of monoclonal antibodies directed against reovirus proteins. Virology 187:760-776. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

