Intramembrane proteolysis promotes trafficking of hepatitis C virus core protein to lipid droplets
- PMID: 12145199
- PMCID: PMC126158
- DOI: 10.1093/emboj/cdf414
Intramembrane proteolysis promotes trafficking of hepatitis C virus core protein to lipid droplets
Abstract
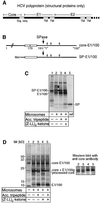
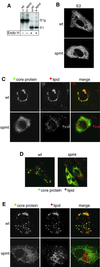
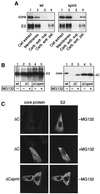
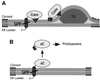
Hepatitis C virus (HCV) is the major causative pathogen associated with liver cirrhosis and hepatocellular carcinoma. The virus has a positive-sense RNA genome encoding a single polyprotein with the virion components located in the N-terminal portion. During biosynthesis of the polyprotein, an internal signal sequence between the core protein and the envelope protein E1 targets the nascent polypeptide to the endoplasmic reticulum (ER) membrane for translocation of E1 into the ER. Following membrane insertion, the signal sequence is cleaved from E1 by signal peptidase. Here we provide evidence that after cleavage by signal peptidase, the signal peptide is further processed by the intramembrane-cleaving protease SPP that promotes the release of core protein from the ER membrane. Core protein is then free for subsequent trafficking to lipid droplets. This study represents an example of a potential role for intramembrane proteolysis in the maturation of a viral protein.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

