Myc and Mad bHLHZ domains possess identical DNA-binding specificities but only partially overlapping functions in vivo
- PMID: 12149476
- PMCID: PMC124931
- DOI: 10.1073/pnas.162369299
Myc and Mad bHLHZ domains possess identical DNA-binding specificities but only partially overlapping functions in vivo
Abstract
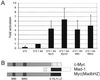
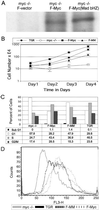
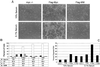
The opposing transcriptional activities of the basic-helix-loop-helix-leucine zipper proteins Myc and Mad, taken together with information related to their expression patterns and biological effects, have led to a model of the Myc/Max/Mad network in which Myc and Mad proteins function as antagonists. This antagonism is presumed to operate at the level of genes targeted by these complexes, where Myc:Max activates and Mad:Max represses expression of the same set of genes. However, a detailed analysis of the DNA-binding preferences for Mad proteins has not been performed. Furthermore, the model does not address the findings that Myc:Max indirectly represses transcription of several regulatory genes. To examine these issues relating to DNA-binding specificity and biological responses, we have determined the DNA-binding preferences of Mad1 by using selection and amplification of randomized oligonucleotides and demonstrated that its intrinsic specificity is identical with that of c-Myc. We have also used a chimeric Myc protein, containing a substitution of the entire Mad basic-helix-loop-helix-leucine zipper motif, and shown that it can reproduce the growth-promoting activities of Myc, but not its apoptotic function. Our results suggest that Myc and Mad, although possessing identical in vitro DNA-binding specificities, do not have an identical set of target genes in vivo, and that apoptosis is one biological outcome in which the transcriptional effects of Myc are not directly antagonized by those of Mad.
Figures

Similar articles
-
Mmip1: a novel leucine zipper protein that reverses the suppressive effects of Mad family members on c-myc.Oncogene. 1998 Mar 5;16(9):1149-59. doi: 10.1038/sj.onc.1201634. Oncogene. 1998. PMID: 9528857
-
Design and properties of a Myc derivative that efficiently homodimerizes.Oncogene. 1998 Nov 12;17(19):2463-72. doi: 10.1038/sj.onc.1202199. Oncogene. 1998. PMID: 9824157
-
Analysis of the Myc and Max interaction specificity with lambda repressor-HLH domain fusions.J Mol Biol. 1995 May 5;248(3):541-50. doi: 10.1006/jmbi.1995.0241. J Mol Biol. 1995. PMID: 7752223
-
The basic region/helix-loop-helix/leucine zipper domain of Myc proto-oncoproteins: function and regulation.Oncogene. 1999 May 13;18(19):2955-66. doi: 10.1038/sj.onc.1202750. Oncogene. 1999. PMID: 10378692 Review.
-
The Max transcription factor network: involvement of Mad in differentiation and an approach to identification of target genes.Cold Spring Harb Symp Quant Biol. 1994;59:109-16. doi: 10.1101/sqb.1994.059.01.014. Cold Spring Harb Symp Quant Biol. 1994. PMID: 7587059 Review.
Cited by
-
cMyc/miR-125b-5p signalling determines sensitivity to bortezomib in preclinical model of cutaneous T-cell lymphomas.PLoS One. 2013;8(3):e59390. doi: 10.1371/journal.pone.0059390. Epub 2013 Mar 19. PLoS One. 2013. PMID: 23527180 Free PMC article.
-
Genome recognition by MYC.Cold Spring Harb Perspect Med. 2014 Feb 1;4(2):a014191. doi: 10.1101/cshperspect.a014191. Cold Spring Harb Perspect Med. 2014. PMID: 24492846 Free PMC article. Review.
-
Myc's broad reach.Genes Dev. 2008 Oct 15;22(20):2755-66. doi: 10.1101/gad.1712408. Genes Dev. 2008. PMID: 18923074 Free PMC article. Review.
-
Genomic binding by the Drosophila Myc, Max, Mad/Mnt transcription factor network.Genes Dev. 2003 May 1;17(9):1101-14. doi: 10.1101/gad.1066903. Epub 2003 Apr 14. Genes Dev. 2003. PMID: 12695332 Free PMC article.
-
DNA methylation analyses of urothelial carcinoma reveal distinct epigenetic subtypes and an association between gene copy number and methylation status.Epigenetics. 2012 Aug;7(8):858-67. doi: 10.4161/epi.20837. Epub 2012 Jun 18. Epigenetics. 2012. PMID: 22705924 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

