Genetically stable picornavirus expression vectors with recombinant internal ribosomal entry sites
- PMID: 12163617
- PMCID: PMC136971
- DOI: 10.1128/jvi.76.17.8966-8972.2002
Genetically stable picornavirus expression vectors with recombinant internal ribosomal entry sites
Abstract
In many respects, picornaviruses are well suited for their proposed use as immunization vectors. However, their inherent genetic instability hinders application for prophylactic purposes. We demonstrate the improved expression and stability of a heterologous insert through a novel vector design strategy that partially replaces noncoding regulatory sequences with coding sequences for foreign gene products.
Figures

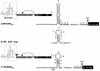

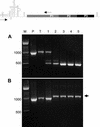


Similar articles
-
Structural determinants of insert retention of poliovirus expression vectors with recombinant IRES elements.Virology. 2003 Jul 5;311(2):241-53. doi: 10.1016/s0042-6822(03)00191-0. Virology. 2003. PMID: 12842615
-
Activity of a type 1 picornavirus internal ribosomal entry site is determined by sequences within the 3' nontranslated region.Proc Natl Acad Sci U S A. 2003 Dec 9;100(25):15125-30. doi: 10.1073/pnas.2436464100. Epub 2003 Nov 26. Proc Natl Acad Sci U S A. 2003. PMID: 14645707 Free PMC article.
-
Retroviral vectors containing putative internal ribosome entry sites: development of a polycistronic gene transfer system and applications to human gene therapy.Nucleic Acids Res. 1992 Mar 25;20(6):1293-9. doi: 10.1093/nar/20.6.1293. Nucleic Acids Res. 1992. PMID: 1313966 Free PMC article.
-
Internal initiation: IRES elements of picornaviruses and hepatitis c virus.Virus Res. 2006 Jul;119(1):2-15. doi: 10.1016/j.virusres.2005.11.003. Epub 2005 Dec 27. Virus Res. 2006. PMID: 16377015 Review.
-
Recombination among picornaviruses.Rev Med Virol. 2010 Sep;20(5):327-37. doi: 10.1002/rmv.660. Rev Med Virol. 2010. PMID: 20632373 Review.
Cited by
-
Packaging of porcine reproductive and respiratory syndrome virus replicon RNA by a stable cell line expressing its nucleocapsid protein.J Microbiol. 2011 Jun;49(3):516-23. doi: 10.1007/s12275-011-1280-1. Epub 2011 Jun 30. J Microbiol. 2011. PMID: 21717343 Free PMC article.
-
Therapeutic Use of Native and Recombinant Enteroviruses.Viruses. 2016 Feb 23;8(3):57. doi: 10.3390/v8030057. Viruses. 2016. PMID: 26907330 Free PMC article. Review.
-
Genetically stable poliovirus vectors activate dendritic cells and prime antitumor CD8 T cell immunity.Nat Commun. 2020 Jan 27;11(1):524. doi: 10.1038/s41467-019-13939-z. Nat Commun. 2020. PMID: 31988324 Free PMC article.
-
Genetic determinants of cell type-specific poliovirus propagation in HEK 293 cells.J Virol. 2005 May;79(10):6281-90. doi: 10.1128/JVI.79.10.6281-6290.2005. J Virol. 2005. PMID: 15858012 Free PMC article.
-
Poliovirus receptor CD155-targeted oncolysis of glioma.Neuro Oncol. 2004 Jul;6(3):208-17. doi: 10.1215/S1152851703000577. Neuro Oncol. 2004. PMID: 15279713 Free PMC article.
References
-
- Agol, V. I., E. V. Pilipenko, and O. R. Slobodskaya. 1996. Modification of translational control elements as a new approach to design of attenuated picornavirus strains. J. Biotechnol. 44:119-128. - PubMed
-
- Almond, J. W., D. Stone, K. Burke, M. A. Skinner, A. J. Macadam, D. Wood, M. Ferguson, and P. D. Minor. 1993. Approaches to the construction of new candidate poliovirus type 3 vaccine strains. Dev. Biol. Stand. 78:161-169. - PubMed
-
- Andino, R., D. S. D. Silvera, P. L. Suggett, C. J. Achacoso, C. J. Miller, D. Baltimore, and M. B. Feinberg. 1994. Engineering poliovirus as a vaccine vector for the expression of diverse antigens. Science 265:1448-1451. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

