The core of the polycomb repressive complex is compositionally and functionally conserved in flies and humans
- PMID: 12167701
- PMCID: PMC134016
- DOI: 10.1128/MCB.22.17.6070-6078.2002
The core of the polycomb repressive complex is compositionally and functionally conserved in flies and humans
Abstract
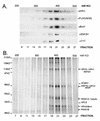
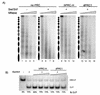
The Polycomb group (PcG) genes are required to maintain homeotic genes in a silenced state during development in drosophila and mammals and are thought to form several distinct silencing complexes that maintain homeotic gene repression during development. Mutations in the PcG genes result in developmental defects and have been implicated in human cancer. Although some PcG protein domains are conserved between flies and humans, substantial regions of several PcG proteins are divergent and humans contain multiple versions of each PcG gene. To determine the effects of these changes on the composition and function of a PcG complex, we have purified a human Polycomb repressive complex from HeLa cells (hPRC-H) that contains homologues of PcG proteins found in drosophila embryonic PRC1 (dPRC1). hPRC-H was found to have fewer components than dPRC1, retaining the PcG core proteins of dPRC1 but lacking most non-PcG proteins. Preparations of hPRC-H contained either two or three different homologues of most of the core PcG proteins, including a new Ph homologue we have named HPH3. Despite differences in composition, dPRC1 and hPRC-H have similar functions: hPRC-H is able to efficiently block remodeling of nucleosomal arrays through a mechanism that does not block the ability of nucleases to access and cleave the arrays.
Figures



References
-
- Akasaka, T., M. Kanno, R. Balling, M. A. Mieza, M. Taniguchi, and H. Koseki. 1996. A role for mel-18, a Polycomb group-related vertebrate gene, during the anteroposterior specification of the axial skeleton. Development 122:1513-1522. - PubMed
-
- Akasaka, T., M. van Lohuizen, N. van der Lugt, Y. Mizutani-Koseki, M. Kanno, M. Taniguchi, M. Vidal, M. Alkema, A. Berns, and H. Koseki. 2001. Mice doubly deficient for the Polycomb group genes Mel18 and Bmi1 reveal synergy and requirement for maintenance but not initiation of Hox gene expression. Development 128:1587-1597. - PubMed
-
- Alkema, M. J., M. Bronk, E. Verhoeven, A. Otte, L. J. van't Veer, A. Berns, and M. van Lohuizen. 1997. Identification of Bmi1-interacting proteins as constituents of a multimeric mammalian Polycomb complex. Genes Dev. 11:226-240. - PubMed
-
- Bardos, J. I., A. J. Saurin, C. Tissot, E. Duprez, and P. S. Freemont. 2000. HPC3 is a new human polycomb orthologue that interacts and associates with RING1 and Bmi1 and has transcriptional repression properties. J. Biol. Chem. 275:28785-28792. - PubMed
-
- Berger, J., H. Kurahashi, Y. Takihara, K. Shimada, H. W. Brock, and F. Randazzo. 1999. The human homolog of Sex comb on midleg (SCMH1) maps to chromosome 1p34. Gene 237:185-191. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
