Identification of a DtxR-regulated operon that is essential for siderophore-dependent iron uptake in Corynebacterium diphtheriae
- PMID: 12169610
- PMCID: PMC135300
- DOI: 10.1128/JB.184.17.4846-4856.2002
Identification of a DtxR-regulated operon that is essential for siderophore-dependent iron uptake in Corynebacterium diphtheriae
Abstract
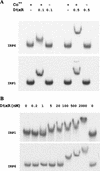
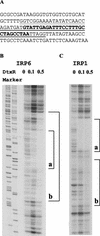
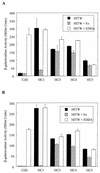
The diphtheria toxin repressor (DtxR) uses Fe(2+) as a corepressor and inhibits transcription from iron-regulated promoters (IRPs) in Corynebacterium diphtheriae. A new IRP, designated IRP6, was cloned from C. diphtheriae by a SELEX-like procedure. DtxR bound to IRP6 in vitro only in the presence of appropriate divalent metal ions, and repression of IRP6 by DtxR in an Escherichia coli system was iron dependent. The open reading frames (ORFs) downstream from IRP6 and previously described promoter IRP1 were found to encode proteins homologous to components of ATP-binding cassette (ABC) transport systems involved in high-affinity iron uptake in other bacteria. IRP1 and IRP6 were repressed under high-iron conditions in wild-type C. diphtheriae C7(beta), but they were expressed constitutively in C7(beta) mutant strains HC1, HC3, HC4, and HC5, which were shown previously to be defective in corynebactin-dependent iron uptake. A clone of the wild-type irp6 operon (pCM6ABC) complemented the constitutive corynebactin production phenotype of HC1, HC4, and HC5 but not of HC3, whereas a clone of the wild-type irp1 operon failed to complement any of these strains. Complementation by subclones of pCM6ABC demonstrated that mutant alleles of irp6A, irp6C, and irp6B were responsible for the phenotypes of HC1, HC4, and HC5, respectively. The irp6A allele in HC1 and the irp6B allele in HC5 encoded single amino acid substitutions in their predicted protein products, and the irp6C allele in HC4 caused premature chain termination of its predicted protein product. Strain HC3 was found to have a chain-terminating mutation in dtxR in addition to a missense mutation in its irp6B allele. These findings demonstrated that the irp6 operon in C. diphtheriae encodes a putative ABC transporter, that specific mutant alleles of irp6A, irp6B, and irp6C are associated with defects in corynebactin-dependent iron uptake, and that complementation of these mutant alleles restores repression of corynebactin production under high-iron growth conditions, most likely as a consequence of restoring siderophore-dependent iron uptake mediated by the irp6 operon.
Figures


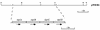
Similar articles
-
Identification and characterization of three new promoter/operators from Corynebacterium diphtheriae that are regulated by the diphtheria toxin repressor (DtxR) and iron.Infect Immun. 1997 Oct;65(10):4273-80. doi: 10.1128/iai.65.10.4273-4280.1997. Infect Immun. 1997. PMID: 9317037 Free PMC article.
-
Analysis of the Corynebacterium diphtheriae DtxR regulon: identification of a putative siderophore synthesis and transport system that is similar to the Yersinia high-pathogenicity island-encoded yersiniabactin synthesis and uptake system.J Bacteriol. 2003 Dec;185(23):6826-40. doi: 10.1128/JB.185.23.6826-6840.2003. J Bacteriol. 2003. PMID: 14617647 Free PMC article.
-
Characterization of an iron-dependent regulatory protein (IdeR) of Mycobacterium tuberculosis as a functional homolog of the diphtheria toxin repressor (DtxR) from Corynebacterium diphtheriae.Infect Immun. 1995 Nov;63(11):4284-9. doi: 10.1128/iai.63.11.4284-4289.1995. Infect Immun. 1995. PMID: 7591059 Free PMC article.
-
Iron, DtxR, and the regulation of diphtheria toxin expression.Mol Microbiol. 1994 Oct;14(2):191-7. doi: 10.1111/j.1365-2958.1994.tb01280.x. Mol Microbiol. 1994. PMID: 7830565 Review.
-
Biology and molecular epidemiology of diphtheria toxin and the tox gene.J Infect Dis. 2000 Feb;181 Suppl 1:S156-67. doi: 10.1086/315554. J Infect Dis. 2000. PMID: 10657208 Review.
Cited by
-
Genomic Insights Into Plant-Growth-Promoting Potentialities of the Genus Frankia.Front Microbiol. 2019 Jul 4;10:1457. doi: 10.3389/fmicb.2019.01457. eCollection 2019. Front Microbiol. 2019. PMID: 31333602 Free PMC article.
-
Analysis of a DtxR-like metalloregulatory protein, MntR, from Corynebacterium diphtheriae that controls expression of an ABC metal transporter by an Mn(2+)-dependent mechanism.J Bacteriol. 2002 Dec;184(24):6882-92. doi: 10.1128/JB.184.24.6882-6892.2002. J Bacteriol. 2002. PMID: 12446639 Free PMC article.
-
Analysis of a heme-dependent signal transduction system in Corynebacterium diphtheriae: deletion of the chrAS genes results in heme sensitivity and diminished heme-dependent activation of the hmuO promoter.Infect Immun. 2005 Nov;73(11):7406-12. doi: 10.1128/IAI.73.11.7406-7412.2005. Infect Immun. 2005. PMID: 16239540 Free PMC article.
-
Molecular characterization of diphtheria toxin repressor (dtxR) genes present in nontoxigenic Corynebacterium diphtheriae strains isolated in the United Kingdom.J Clin Microbiol. 2005 Jan;43(1):223-8. doi: 10.1128/JCM.43.1.223-228.2005. J Clin Microbiol. 2005. PMID: 15634975 Free PMC article.
-
Pan-genomic analysis of Corynebacterium amycolatum gives insights into molecular mechanisms underpinning the transition to a pathogenic phenotype.Front Microbiol. 2022 Nov 16;13:1011578. doi: 10.3389/fmicb.2022.1011578. eCollection 2022. Front Microbiol. 2022. PMID: 36466658 Free PMC article.
References
-
- Budzikiewicz, H., A. Bössenkamp, K. Taraz, A. Pandey, and J.-M. Meyer. 1997. Corynebactin, a cyclic catecholate siderophore from Corynebacterium glutamicum ATCC 14067 (Brevibacterium sp. DSM 20411). Z. Naturforsch. Sect. C 52:551-554.
-
- Drazek, E. S., C. A. Hammack, Sr., and M. P. Schmitt. 2000. Corynebacterium diphtheriae genes required for acquisition of iron from haemin and haemoglobin are homologous to ABC haemin transporters. Mol. Microbiol. 36:68-84. - PubMed
-
- Feese, M. D., E. Pohl, R. K. Holmes, and W. G. J. Hol. 2001. Iron-dependent regulators, p. 850-863. In A. Messerschmidt, R. Huber, T. Poulos, and K. Wieghardt (ed.), Handbook of metalloproteins. John Wiley & Sons, Ltd., Chichester, United Kingdom.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

