Counting CAG repeats in the Huntington's disease gene by restriction endonuclease EcoP15I cleavage
- PMID: 12177311
- PMCID: PMC134256
- DOI: 10.1093/nar/gnf082
Counting CAG repeats in the Huntington's disease gene by restriction endonuclease EcoP15I cleavage
Abstract
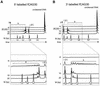
Huntington's disease (HD) is a progressive neurodegenerative disorder with autosomal-dominant inheritance. The disease is caused by a CAG trinucleotide repeat expansion located in the first exon of the HD gene. The CAG repeat is highly polymorphic and varies from 6 to 37 repeats on chromosomes of unaffected individuals and from more than 30 to 180 repeats on chromosomes of HD patients. In this study, we show that the number of CAG repeats in the HD gene can be determined by restriction of the DNA with the endonuclease EcoP15I and subsequent analysis of the restriction fragment pattern by electrophoresis through non-denaturing polyacrylamide gels using the ALFexpress DNA Analysis System. CAG repeat numbers in the normal (30 and 35 repeats) as well as in the pathological range (81 repeats) could be accurately counted using this assay. Our results suggest that this high-resolution method can be used for the exact length determination of CAG repeats in HD genes as well as in genes affected in related CAG repeat disorders.
Figures



Similar articles
-
Autopsy-proven Huntington's disease with 29 trinucleotide repeats.Mov Disord. 2007 Jan;22(1):127-30. doi: 10.1002/mds.21195. Mov Disord. 2007. PMID: 17115386
-
Null alleles at the Huntington disease locus: implications for diagnostics and CAG repeat instability.Genet Test. 2000;4(1):55-60. doi: 10.1089/109065700316480. Genet Test. 2000. PMID: 10794362
-
Searching for mutation in the JPH3, ATN1 and TBP genes in Polish patients suspected of Huntington's disease and without mutation in the IT15 gene.Neurol Neurochir Pol. 2008 May-Jun;42(3):203-9. Neurol Neurochir Pol. 2008. PMID: 18651325
-
Genetics and neuropathology of Huntington's disease.Int Rev Neurobiol. 2011;98:325-72. doi: 10.1016/B978-0-12-381328-2.00014-6. Int Rev Neurobiol. 2011. PMID: 21907094 Free PMC article. Review.
-
[Huntington's disease--advances in gene mapping].Nihon Rinsho. 1993 Sep;51(9):2481-7. Nihon Rinsho. 1993. PMID: 8411732 Review. Japanese.
Cited by
-
Identification of restriction endonucleases sensitive to 5-cytosine methylation at non-CpG sites, including expanded (CAG)n/(CTG)n repeats.Epigenetics. 2011 Apr;6(4):416-20. doi: 10.4161/epi.6.4.14953. Epub 2011 Apr 1. Epigenetics. 2011. PMID: 21364324 Free PMC article.
-
Role of Dynein Axonemal Heavy Chain 6 Gene Expression as a Possible Biomarker for Huntington's Disease: a Translational Study.J Mol Neurosci. 2017 Dec;63(3-4):342-348. doi: 10.1007/s12031-017-0984-z. Epub 2017 Oct 10. J Mol Neurosci. 2017. PMID: 29019003
-
Structural insights into the assembly and shape of Type III restriction-modification (R-M) EcoP15I complex by small-angle X-ray scattering.J Mol Biol. 2012 Jul 20;420(4-5):261-8. doi: 10.1016/j.jmb.2012.04.026. Epub 2012 May 2. J Mol Biol. 2012. PMID: 22560991 Free PMC article.
-
An Evolutionary Perspective on the Impact of Genomic Copy Number Variation on Human Health.J Mol Evol. 2020 Jan;88(1):104-119. doi: 10.1007/s00239-019-09911-6. Epub 2019 Sep 14. J Mol Evol. 2020. PMID: 31522275 Review.
-
Antagonistic pleiotropy as a widespread mechanism for the maintenance of polymorphic disease alleles.BMC Med Genet. 2011 Dec 12;12:160. doi: 10.1186/1471-2350-12-160. BMC Med Genet. 2011. PMID: 22151998 Free PMC article.
References
-
- Wanker E.E. (2000) Protein aggregation and pathogenesis of Huntington’s disease: mechanisms and correlations. Biol. Chem., 381, 937–942. - PubMed
-
- Gusella J.F. and MacDonald,M.E. (2000) Molecular genetics: unmasking polyglutamine triggers in neurodegenerative disease. Nature Rev. Neurosci., 1, 109–115. - PubMed
-
- Scherzinger E., Sittler,A., Schweiger,K., Heiser,V., Lurz,R., Hasenbank,R., Bates,G.P., Lehrach,H. and Wanker,E.E. (1999) Self-assembly of polyglutamine-containing huntingtin fragments into amyloid-like fibrils: implications for Huntington’s disease pathology. Proc. Natl Acad. Sci. USA, 96, 4604–4609. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

