Histone H3 variants specify modes of chromatin assembly
- PMID: 12177448
- PMCID: PMC139911
- DOI: 10.1073/pnas.172403699
Histone H3 variants specify modes of chromatin assembly
Abstract
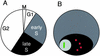
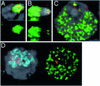
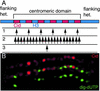
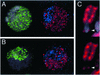
Histone variants have been known for 30 years, but their functions and the mechanism of their deposition are still largely unknown. Drosophila has three versions of histone H3. H3 packages the bulk genome, H3.3 marks active chromatin and may be essential for gene regulation, and Cid is the characteristic structural component of centromeric chromatin. We have characterized the properties of these histones by using a Drosophila cell-line system that allows precise analysis of both DNA replication and histone deposition. The deposition of H3 is restricted to replicating DNA. In striking contrast, H3.3 and Cid deposit throughout the cell cycle. Deposition of H3.3 occurs without any corresponding DNA replication. To confirm that the deposition of Cid is also replication-independent (RI), we examined centromere replication in cultured cells and neuroblasts. We found that centromeres replicate out of phase with heterochromatin and display replication patterns that may limit H3 deposition. This confirms that both variants undergo RI deposition, but at different locations in the nucleus. How variant histones accomplish RI deposition is unknown, and raises basic questions about the stability of nucleosomes, the machinery that accomplishes nucleosome assembly, and the functional organization of the nucleus. The different in vivo properties of H3, H3.3, and Cid set the stage for identifying the mechanisms by which they are differentially targeted. Here we suggest that local effects of "open" chromatin and broader effects of nuclear organization help to guide the two different H3 variants to their target sites.
Figures

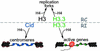
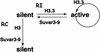

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

