Normalization and analysis of DNA microarray data by self-consistency and local regression
- PMID: 12184811
- PMCID: PMC126242
- DOI: 10.1186/gb-2002-3-7-research0037
Normalization and analysis of DNA microarray data by self-consistency and local regression
Abstract
Background: With the advent of DNA hybridization microarrays comes the remarkable ability, in principle, to simultaneously monitor the expression levels of thousands of genes. The quantitative comparison of two or more microarrays can reveal, for example, the distinct patterns of gene expression that define different cellular phenotypes or the genes induced in the cellular response to insult or changing environmental conditions. Normalization of the measured intensities is a prerequisite of such comparisons, and indeed, of any statistical analysis, yet insufficient attention has been paid to its systematic study. The most straightforward normalization techniques in use rest on the implicit assumption of linear response between true expression level and output intensity. We find that these assumptions are not generally met, and that these simple methods can be improved.
Results: We have developed a robust semi-parametric normalization technique based on the assumption that the large majority of genes will not have their relative expression levels changed from one treatment group to the next, and on the assumption that departures of the response from linearity are small and slowly varying. We use local regression to estimate the normalized expression levels as well as the expression level-dependent error variance.
Conclusions: We illustrate the use of this technique in a comparison of the expression profiles of cultured rat mesothelioma cells under control and under treatment with potassium bromate, validated using quantitative PCR on a selected set of genes. We tested the method using data simulated under various error models and find that it performs well.
Figures

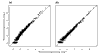
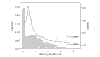

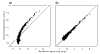
Similar articles
-
Evaluating different methods of microarray data normalization.BMC Bioinformatics. 2006 Oct 23;7:469. doi: 10.1186/1471-2105-7-469. BMC Bioinformatics. 2006. PMID: 17059609 Free PMC article.
-
Evaluation of normalization methods for microarray data.BMC Bioinformatics. 2003 Sep 2;4:33. doi: 10.1186/1471-2105-4-33. Epub 2003 Sep 2. BMC Bioinformatics. 2003. PMID: 12950995 Free PMC article.
-
SED, a normalization free method for DNA microarray data analysis.BMC Bioinformatics. 2004 Sep 2;5:121. doi: 10.1186/1471-2105-5-121. BMC Bioinformatics. 2004. PMID: 15345033 Free PMC article.
-
Normalization of microarray data: single-labeled and dual-labeled arrays.Mol Cells. 2006 Dec 31;22(3):254-61. Mol Cells. 2006. PMID: 17202852 Review.
-
Statistical analysis of microarray data.Addict Biol. 2005 Mar;10(1):23-35. doi: 10.1080/13556210412331327795. Addict Biol. 2005. PMID: 15849016 Review.
Cited by
-
Temperature effects on DNA chip experiments from surface plasmon resonance imaging: isotherms and melting curves.Biophys J. 2007 Feb 1;92(3):935-46. doi: 10.1529/biophysj.106.097790. Epub 2006 Nov 3. Biophys J. 2007. PMID: 17085497 Free PMC article.
-
Profound influence of microarray scanner characteristics on gene expression ratios: analysis and procedure for correction.BMC Genomics. 2004 Feb 3;5(1):10. doi: 10.1186/1471-2164-5-10. BMC Genomics. 2004. PMID: 15018648 Free PMC article.
-
Accurate and reliable cancer classification based on probabilistic inference of pathway activity.PLoS One. 2009 Dec 7;4(12):e8161. doi: 10.1371/journal.pone.0008161. PLoS One. 2009. PMID: 19997592 Free PMC article.
-
Pathway level analysis of gene expression using singular value decomposition.BMC Bioinformatics. 2005 Sep 12;6:225. doi: 10.1186/1471-2105-6-225. BMC Bioinformatics. 2005. PMID: 16156896 Free PMC article.
-
Identification of diagnostic subnetwork markers for cancer in human protein-protein interaction network.BMC Bioinformatics. 2010 Oct 7;11 Suppl 6(Suppl 6):S8. doi: 10.1186/1471-2105-11-S6-S8. BMC Bioinformatics. 2010. PMID: 20946619 Free PMC article.
References
-
- Fodor SP, Rava RP, Huang XC, Pease AC, Holmes CP, Adams CL. Multiplexed biochemical assays with biological chips. Nature. 1993;364:555–556. - PubMed
-
- Schena M, Shalon D, Davis RW, Brown PO. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science. 1995;270:467–470. - PubMed
-
- DeRisi J, Penland L, Brown PO, Bittner ML, Meltzer PP, Ray M, Chen Y, Su YA, Trent JM. Use of a cDNA microarray to analyze gene expression patterns in human cancer. Nat Genet. 1996;14:457–460. - PubMed
-
- Lockhart DJ, Dong H, Byrne MC, Follettie MT, Gallo MV, Chee MS, Mittmann M, Wang C, Kobayashi M, Horton H, Brown EL. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat Biotechnol. 1996;14:1675–1680. - PubMed
-
- DeRisi JL, Iyer VR, Brown PO. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science. 1997;278:680–686. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

