Structural insights into peptide bond formation
- PMID: 12185246
- PMCID: PMC129327
- DOI: 10.1073/pnas.172404099
Structural insights into peptide bond formation
Abstract
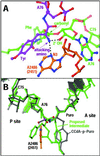
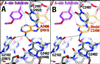
The large ribosomal subunit catalyzes peptide bond formation and will do so by using small aminoacyl- and peptidyl-RNA fragments of tRNA. We have refined at 3-A resolution the structures of both A and P site substrate and product analogues, as well as an intermediate analogue, bound to the Haloarcula marismortui 50S ribosomal subunit. A P site substrate, CCA-Phe-caproic acid-biotin, binds equally to both sites, but in the presence of sparsomycin binds only to the P site. The CCA portions of these analogues are bound identically by either the A or P loop of the 23S rRNA. Combining the separate P and A site substrate complexes into one model reveals interactions that may occur when both are present simultaneously. The alpha-NH(2) group of an aminoacylated fragment in the A site forms one hydrogen bond with the N3 of A2486 (2451) and may form a second hydrogen bond either with the 2' OH of the A-76 ribose in the P site or with the 2' OH of A2486 (2451). These interactions position the alpha amino group adjacent to the carbonyl carbon of esterified P site substrate in an orientation suitable for a nucleophilic attack.
Figures



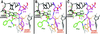
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

