Characterization of RAD51-independent break-induced replication that acts preferentially with short homologous sequences
- PMID: 12192038
- PMCID: PMC135638
- DOI: 10.1128/MCB.22.18.6384-6392.2002
Characterization of RAD51-independent break-induced replication that acts preferentially with short homologous sequences
Abstract
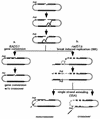
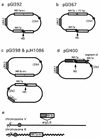
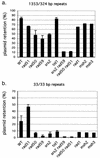
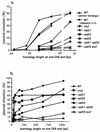
Repair of double-strand breaks by gene conversions between homologous sequences located on different Saccharomyces cerevisiae chromosomes or plasmids requires RAD51. When repair occurs between inverted repeats of the same plasmid, both RAD51-dependent and RAD51-independent repairs are found. Completion of RAD51-independent plasmid repair events requires RAD52, RAD50, RAD59, TID1 (RDH54), and SRS2 and appears to involve break-induced replication coupled to single-strand annealing. Surprisingly, RAD51-independent recombination requires much less homology (30 bp) for strand invasion than does RAD51-dependent repair (approximately 100 bp); in fact, the presence of Rad51p impairs recombination with short homology. The differences between the RAD51- and RAD50/RAD59-dependent pathways account for the distinct ways that two different recombination processes maintain yeast telomeres in the absence of telomerase.
Figures

References
-
- Bai, Y., and L. S. Symington. 1996. A Rad52 homolog is required for RAD51-independent mitotic recombination in Saccharomyces cerevisiae. Genes Dev. 10:2025-2037. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
