Regulation of the premiddle and middle phases of expression of the NDT80 gene during sporulation of Saccharomyces cerevisiae
- PMID: 12192041
- PMCID: PMC135636
- DOI: 10.1128/MCB.22.18.6417-6429.2002
Regulation of the premiddle and middle phases of expression of the NDT80 gene during sporulation of Saccharomyces cerevisiae
Abstract
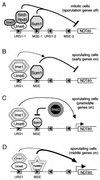
The NDT80 gene of Saccharomyces cerevisiae, which encodes a global activator of transcription of middle sporulation-specific genes, is first expressed after the activation of early meiotic genes but prior to activation of middle sporulation-specific genes. Both upstream repression sequence 1 (URS1) and mid-sporulation element (MSE) sites are present in the promoter region of the NDT80 gene; these elements have been shown previously to contribute to the regulation of expression of early and middle sporulation-specific genes, respectively, by mediating repression in growing cells and activation at specific times during sporulation. In this study, we have shown that the overlapping windows of URS1- and MSE-mediated repression and activation are responsible for the distinctive premiddle expression pattern of the NDT80 gene. Our data suggest that a Sum1-associated repression complex bound at the NDT80 MSE sites prevents Ime1 tethered at the NDT80 URS1 sites from activating transcription of the NDT80 gene at the time that Ime1-dependent activation of early URS1-regulated meiotic genes is occurring. We propose that a decrease in the efficiency of Sum1-mediated repression as cells progress through the early events of the sporulation program allows the previously inactive Ime1 tethered at the URS1(NDT80) sites to promote a low level of expression of the NDT80 gene. This initial phase of URS1-dependent NDT80 expression is followed by Ndt80-dependent upregulation of its own expression, which requires the MSE(NDT80) sites and occurs concomitantly with Ndt80-dependent activation of a set of middle MSE-regulated sporulation-specific genes. Mutation of IME2 prevents expression of NDT80 in sporulating cells. We show in this study that NDT80 is expressed and that middle genes are activated in cells of an Deltaime2/Deltaime2 Deltasum1/Deltasum1 strain in sporulation medium. This suggests that Ime2 activates expression of NDT80 by eliminating Sum1-mediated repression.
Figures

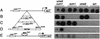





References
-
- Barral, Y., S. Jentsch, and C. Mann. 1995. G1 cyclin turnover and nutrient uptake are controlled by a common pathway in yeast. Genes Dev. 9:399-409. - PubMed
-
- Brachmann, C. B., J. M. Sherman, S. E. Devine, E. E. Cameron, L. Pillus, and J. D. Boeke. 1995. The SIR2 gene family, conserved from bacteria to humans, functions in silencing, cell cycle progression, and chromosome stability. Genes Dev. 9:2888-2902. - PubMed
-
- Chu, S., and I. Herskowitz. 1998. Gametogenesis in yeast is regulated by a transcriptional cascade dependent on Ndt80. Mol. Cell 1:685-696. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
