3' end processing of Drosophila melanogaster histone pre-mRNAs: requirement for phosphorylated Drosophila stem-loop binding protein and coevolution of the histone pre-mRNA processing system
- PMID: 12192062
- PMCID: PMC135633
- DOI: 10.1128/MCB.22.18.6648-6660.2002
3' end processing of Drosophila melanogaster histone pre-mRNAs: requirement for phosphorylated Drosophila stem-loop binding protein and coevolution of the histone pre-mRNA processing system
Abstract
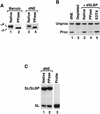
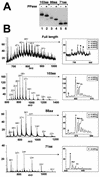
Synthetic pre-mRNAs containing the processing signals encoded by Drosophila melanogaster histone genes undergo efficient and faithful endonucleolytic cleavage in nuclear extracts prepared from Drosophila cultured cells and 0- to 13-h-old embryos. Biochemical requirements for the in vitro cleavage are similar to those previously described for the 3' end processing of mammalian histone pre-mRNAs. Drosophila 3' end processing does not require ATP and occurs in the presence of EDTA. However, in contrast to mammalian processing, Drosophila processing generates the final product ending four nucleotides after the stem-loop. Cleavage of the Drosophila substrates is abolished by depleting the extract of the Drosophila stem-loop binding protein (dSLBP), indicating that both dSLBP and the stem-loop structure in histone pre-mRNA are essential components of the processing machinery. Recombinant dSLBP expressed in insect cells by using the baculovirus system efficiently complements the depleted extract. Only the RNA-binding domain plus the 17 amino acids at the C terminus of dSLBP are required for processing. The full-length dSLBP expressed in insect cells is quantitatively phosphorylated on four residues in the C-terminal region. Dephosphorylation of the recombinant dSLBP reduces processing activity. Human and Drosophila SLBPs are not interchangeable and strongly inhibit processing in the heterologous extracts. The RNA-binding domain of the dSLBP does not substitute for the RNA-binding domain of the human SLBP in histone pre-mRNA processing in mammalian extracts. In addition to the stem-loop structure and dSLBP, 3' processing in Drosophila nuclear extracts depends on the presence of a short stretch of purines located ca. 20 nucleotides downstream from the stem, and an Sm-reactive factor, most likely the Drosophila counterpart of vertebrate U7 snRNP.
Figures



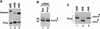
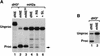

Similar articles
-
Drosophila stem loop binding protein coordinates accumulation of mature histone mRNA with cell cycle progression.Genes Dev. 2001 Jan 15;15(2):173-87. doi: 10.1101/gad.862801. Genes Dev. 2001. PMID: 11157774 Free PMC article.
-
Developmental control of histone mRNA and dSLBP synthesis during Drosophila embryogenesis and the role of dSLBP in histone mRNA 3' end processing in vivo.Mol Cell Biol. 2002 Apr;22(7):2267-82. doi: 10.1128/MCB.22.7.2267-2282.2002. Mol Cell Biol. 2002. PMID: 11884612 Free PMC article.
-
Drosophila stem-loop binding protein intracellular localization is mediated by phosphorylation and is required for cell cycle-regulated histone mRNA expression.Mol Biol Cell. 2004 Mar;15(3):1112-23. doi: 10.1091/mbc.e03-09-0649. Mol Biol Cell. 2004. PMID: 14999087 Free PMC article.
-
Formation of the 3' end of histone mRNA.Gene. 1999 Oct 18;239(1):1-14. doi: 10.1016/s0378-1119(99)00367-4. Gene. 1999. PMID: 10571029 Review.
-
Formation of the 3' end of histone mRNA: getting closer to the end.Gene. 2007 Jul 15;396(2):373-90. doi: 10.1016/j.gene.2007.04.021. Epub 2007 May 4. Gene. 2007. PMID: 17531405 Free PMC article. Review.
Cited by
-
Contribution of protein phosphorylation to binding-induced folding of the SLBP-histone mRNA complex probed by phosphorus-31 NMR.FEBS Open Bio. 2014 Oct 16;4:853-7. doi: 10.1016/j.fob.2014.10.002. eCollection 2014. FEBS Open Bio. 2014. PMID: 25379382 Free PMC article.
-
Structure-specific nucleic acid recognition by L-motifs and their diverse roles in expression and regulation of the genome.Biochim Biophys Acta. 2015 Jun;1849(6):677-87. doi: 10.1016/j.bbagrm.2015.02.006. Epub 2015 Mar 4. Biochim Biophys Acta. 2015. PMID: 25748361 Free PMC article. Review.
-
A region of SLBP outside the mRNA-processing domain is essential for deposition of histone mRNA into the Drosophila egg.J Cell Sci. 2021 Feb 11;134(3):jcs251728. doi: 10.1242/jcs.251728. J Cell Sci. 2021. PMID: 33408246 Free PMC article.
-
3'-End processing of histone pre-mRNAs in Drosophila: U7 snRNP is associated with FLASH and polyadenylation factors.RNA. 2013 Dec;19(12):1726-44. doi: 10.1261/rna.040360.113. Epub 2013 Oct 21. RNA. 2013. PMID: 24145821 Free PMC article.
-
Phosphorylation of threonine 61 by cyclin a/Cdk1 triggers degradation of stem-loop binding protein at the end of S phase.Mol Cell Biol. 2008 Jul;28(14):4469-79. doi: 10.1128/MCB.01416-07. Epub 2008 May 19. Mol Cell Biol. 2008. PMID: 18490441 Free PMC article.
References
-
- Albig, W., and D. Doenecke. 1997. The human histone gene cluster at the D6S105 locus. Hum. Genet. 101:284-294. - PubMed
-
- Albig, W., P. Kioschis, A. Poustka, K. Meergans, and D. Doenecke. 1997. Human histone gene organization: nonregular arrangement within a large cluster. Genomics 40:314-322. - PubMed
-
- Birnstiel, M. L., and F. J. Schaufele. 1988. Structure and function of minor snRNPs, p. 155-182. In M. L. Birnstiel (ed.), Structure and function of major and minor small ribonucleoprotein particles. Springer-Verlag, Berlin, Germany.
-
- Bond, U. M., T. A. Yario, and J. A. Steitz. 1991. Multiple processing-defective mutations in a mammalian histone premessenger RNA are suppressed by compensatory changes in U7 RNA both in vivo and in vitro. Genes Dev. 5:1709-1722. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
