Crystal structures of a meta-cleavage product hydrolase from Pseudomonas fluorescens IP01 (CumD) complexed with cleavage products
- PMID: 12192074
- PMCID: PMC2373588
- DOI: 10.1110/ps.0209602
Crystal structures of a meta-cleavage product hydrolase from Pseudomonas fluorescens IP01 (CumD) complexed with cleavage products
Abstract
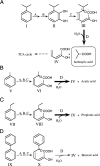
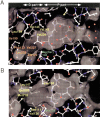
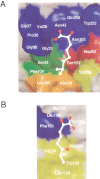
2-Hydroxy-6-oxo-7-methylocta-2,4-dienoate hydrolase (CumD) from Pseudomonas fluorescens IP01 hydrolyzes a meta-cleavage product generated in the cumene (isopropylbenzene) degradation pathway. The crystal structures of the inactive S103A mutant of the CumD enzyme complexed with isobutyrate and acetate ions were determined at 1.6 and 2.0 A resolution, respectively. The isobutyrate and acetate ions were located at the same position in the active site, and occupied the site for a part of the hydrolysis product with CumD, which has the key determinant group for the substrate specificity of related hydrolases. One of the oxygen atoms of the carboxyl group of the isobutyrate ion was hydrogen bonded with a water molecule and His252. Another oxygen atom of the carboxyl group was situated in an oxyanion hole formed by the two main-chain N atoms. The isopropyl group of the isobutyric acid was recognized by the side-chains of the hydrophobic residues. The substrate-binding pocket of CumD was long, and the inhibition constants of various organic acids corresponded well to it. In comparison with the structure of BphD from Rhodococcus sp. RHA1, the structural basis for the substrate specificity of related hydrolases, is revealed.
Figures

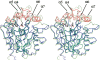

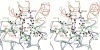
Similar articles
-
Improving the catalytic efficiency of a meta-cleavage product hydrolase (CumD) from Pseudomonas fluorescens IP01.Biochim Biophys Acta. 2006 Jul;1764(7):1159-66. doi: 10.1016/j.bbapap.2006.05.010. Epub 2006 Jun 7. Biochim Biophys Acta. 2006. PMID: 16844437
-
A series of crystal structures of a meta-cleavage product hydrolase from Pseudomonas fluorescens IP01 (CumD) complexed with various cleavage products.Biosci Biotechnol Biochem. 2005 Mar;69(3):491-8. doi: 10.1271/bbb.69.491. Biosci Biotechnol Biochem. 2005. PMID: 15784976
-
Crystal structure of 2-hydroxyl-6-oxo-6-phenylhexa-2,4-dienoic acid (HPDA) hydrolase (BphD enzyme) from the Rhodococcus sp. strain RHA1 of the PCB degradation pathway.J Mol Biol. 2001 Jun 22;309(5):1139-51. doi: 10.1006/jmbi.2001.4737. J Mol Biol. 2001. PMID: 11399084
-
Structure and mechanism of kynureninase.Arch Biochem Biophys. 2014 Feb 15;544:69-74. doi: 10.1016/j.abb.2013.10.020. Epub 2013 Nov 4. Arch Biochem Biophys. 2014. PMID: 24200862 Review.
-
Structure, mechanism, and substrate specificity of kynureninase.Biochim Biophys Acta. 2011 Nov;1814(11):1481-8. doi: 10.1016/j.bbapap.2010.12.003. Epub 2010 Dec 15. Biochim Biophys Acta. 2011. PMID: 21167323 Free PMC article. Review.
Cited by
-
Kinetic and structural insight into the mechanism of BphD, a C-C bond hydrolase from the biphenyl degradation pathway.Biochemistry. 2006 Sep 19;45(37):11071-86. doi: 10.1021/bi0611098. Biochemistry. 2006. PMID: 16964968 Free PMC article.
-
Cloning, expression and characterization of a lipase gene (lip3) from Pseudomonas aeruginosa LST-03.Mol Genet Genomics. 2004 Mar;271(2):189-96. doi: 10.1007/s00438-003-0970-8. Epub 2004 Jan 23. Mol Genet Genomics. 2004. PMID: 14740297
-
Deacylation on the matrix side of the mitochondrial inner membrane regulates cardiolipin remodeling.Mol Biol Cell. 2013 Jun;24(12):2008-20. doi: 10.1091/mbc.E13-03-0121. Epub 2013 May 1. Mol Biol Cell. 2013. PMID: 23637464 Free PMC article.
-
Characterization of a C-C bond hydrolase from Sphingomonas wittichii RW1 with novel specificities towards polychlorinated biphenyl metabolites.J Bacteriol. 2007 Jun;189(11):4038-45. doi: 10.1128/JB.01950-06. Epub 2007 Apr 6. J Bacteriol. 2007. PMID: 17416660 Free PMC article.
-
The lid domain of the MCP hydrolase DxnB2 contributes to the reactivity toward recalcitrant PCB metabolites.Biochemistry. 2013 Aug 20;52(33):5685-5695. doi: 10.1021/bi400774m. Epub 2013 Aug 9. Biochemistry. 2013. PMID: 23879719 Free PMC article.
References
-
- Aoki, H., Kimura, T., Habe, H., Yamane, H., Kodama, T., and Omori, T. 1996. Cloning, nucleotide sequence, and characterization of the genes encoding enzymes involved in the degradation of cumene to 2-hydroxy-6-oxo-7-methylocta-2,4-dienoic acid in Pseudomonas fluorescens IP01.J. Ferment. Bioeng. 81 187–196.
-
- Bedard, D.L. and Haberl, M.L. 1990. Influence of chlorine substitution pattern on the degradation of polychlorinated biphenyls by eight bacterial strains.Microbiol. Ecol. 20 87–102. - PubMed
-
- Bernstein, H.J. 2000. Recent changes to RasMol, recombining the variants.Trends Biochem. Sci. 25 453–455. - PubMed
-
- Blumer, M. and Youngblood, W.W. 1975. Polycyclic aromatic hydrocarbons in soils and recent sediments.Science 188 53–55. - PubMed
-
- Brunger, A.T., Adams, P.D., Clore, G.M., DeLano, W.L., Gros, P., Grosse-Kunstleve, R.W., Jiang, J.-S., Kuszewski, J., Nilges, M., Pannu, N.S., Read, R.J., Rice, L.M., Simonson, T., and Warren, G.L. 1998. Crystallography and NMR system: A new software suite for macromolecular structure determination.Acta Crystallogr. D Biol. Crystallogr. D54 905–921. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

