Structure and function of the BAH-containing domain of Orc1p in epigenetic silencing
- PMID: 12198162
- PMCID: PMC125411
- DOI: 10.1093/emboj/cdf468
Structure and function of the BAH-containing domain of Orc1p in epigenetic silencing
Abstract
The N-terminal domain of the largest subunit of the Saccharomyces cerevisiae origin recognition complex (Orc1p) functions in transcriptional silencing and contains a bromo-adjacent homology (BAH) domain found in some chromatin-associated proteins including Sir3p. The 2.2 A crystal structure of the N-terminal domain of Orc1p revealed a BAH core and a non-conserved helical sub-domain. Mutational analyses demonstrated that the helical sub-domain was necessary and sufficient to bind Sir1p, and critical for targeting Sir1p primarily to the cis-acting E silencers at the HMR and HML silent chromatin domains. In the absence of the BAH domain, approximately 14-20% of cells in a population were silenced at the HML locus. Moreover, the distributions of the Sir2p, Sir3p and Sir4p proteins, while normal, were at levels lower than found in wild-type cells. Thus, in the absence of the Orc1p BAH domain, HML resembled silencing of genes adjacent to telomeres. These data are consistent with the view that the Orc1p-Sir1p interaction at the E silencers ensures stable inheritance of pre-established Sir2p, Sir3p and Sir4p complexes at the silent mating type loci.
Figures


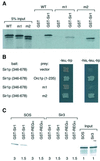

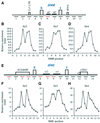
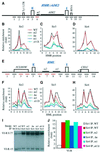

Similar articles
-
Structure of the Sir3 protein bromo adjacent homology (BAH) domain from S. cerevisiae at 1.95 A resolution.Protein Sci. 2006 May;15(5):1182-6. doi: 10.1110/ps.052061006. Protein Sci. 2006. PMID: 16641491 Free PMC article.
-
Structural basis for origin recognition complex 1 protein-silence information regulator 1 protein interaction in epigenetic silencing.Proc Natl Acad Sci U S A. 2005 Jun 14;102(24):8519-24. doi: 10.1073/pnas.0502946102. Epub 2005 Jun 3. Proc Natl Acad Sci U S A. 2005. PMID: 15937111 Free PMC article.
-
The origin recognition complex and Sir4 protein recruit Sir1p to yeast silent chromatin through independent interactions requiring a common Sir1p domain.Mol Cell Biol. 2004 Jan;24(2):774-86. doi: 10.1128/MCB.24.2.774-786.2004. Mol Cell Biol. 2004. PMID: 14701749 Free PMC article.
-
Silent information regulator 3: the Goldilocks of the silencing complex.Genes Dev. 2010 Jan 15;24(2):115-22. doi: 10.1101/gad.1865510. Genes Dev. 2010. PMID: 20080949 Free PMC article. Review.
-
Silent chromatin in yeast: an orchestrated medley featuring Sir3p [corrected].Bioessays. 1998 Jan;20(1):30-40. doi: 10.1002/(SICI)1521-1878(199801)20:1<30::AID-BIES6>3.0.CO;2-W. Bioessays. 1998. PMID: 9504045 Review.
Cited by
-
Nα-acetylated Sir3 stabilizes the conformation of a nucleosome-binding loop in the BAH domain.Nat Struct Mol Biol. 2013 Sep;20(9):1116-8. doi: 10.1038/nsmb.2637. Epub 2013 Aug 11. Nat Struct Mol Biol. 2013. PMID: 23934152
-
A Structural Perspective on Readout of Epigenetic Histone and DNA Methylation Marks.Cold Spring Harb Perspect Biol. 2016 Mar 1;8(3):a018754. doi: 10.1101/cshperspect.a018754. Cold Spring Harb Perspect Biol. 2016. PMID: 26931326 Free PMC article. Review.
-
Dependence of ORC silencing function on NatA-mediated Nalpha acetylation in Saccharomyces cerevisiae.Mol Cell Biol. 2004 Dec;24(23):10300-12. doi: 10.1128/MCB.24.23.10300-10312.2004. Mol Cell Biol. 2004. PMID: 15542839 Free PMC article.
-
Transcriptional silencing functions of the yeast protein Orc1/Sir3 subfunctionalized after gene duplication.Proc Natl Acad Sci U S A. 2010 Nov 9;107(45):19384-9. doi: 10.1073/pnas.1006436107. Epub 2010 Oct 25. Proc Natl Acad Sci U S A. 2010. PMID: 20974972 Free PMC article.
-
Origin recognition complex harbors an intrinsic nucleosome remodeling activity.Proc Natl Acad Sci U S A. 2022 Oct 18;119(42):e2211568119. doi: 10.1073/pnas.2211568119. Epub 2022 Oct 10. Proc Natl Acad Sci U S A. 2022. PMID: 36215487 Free PMC article.
References
-
- Bell S.P. and Stillman,B. (1992) ATP-dependent recognition of eukaryotic origins of DNA replication by a multiprotein complex. Nature, 357, 128–134. - PubMed
-
- Bell S.P., Kobayashi,R. and Stillman,B. (1993) Yeast origin recognition complex functions in transcription silencing and DNA replication. Science, 262, 1844–1849. - PubMed
-
- Bell S.P., Mitchell,J., Leber,J., Kobayashi,R. and Stillman,B. (1995) The multidomain structure of Orc1p reveals similarity to regulators of DNA replication and transcriptional silencing. Cell, 83, 563–568. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

