Human non-synonymous SNPs: server and survey
- PMID: 12202775
- PMCID: PMC137415
- DOI: 10.1093/nar/gkf493
Human non-synonymous SNPs: server and survey
Abstract
Human single nucleotide polymorphisms (SNPs) represent the most frequent type of human population DNA variation. One of the main goals of SNP research is to understand the genetics of the human phenotype variation and especially the genetic basis of human complex diseases. Non-synonymous coding SNPs (nsSNPs) comprise a group of SNPs that, together with SNPs in regulatory regions, are believed to have the highest impact on phenotype. Here we present a World Wide Web server to predict the effect of an nsSNP on protein structure and function. The prediction method enabled analysis of the publicly available SNP database HGVbase, which gave rise to a dataset of nsSNPs with predicted functionality. The dataset was further used to compare the effect of various structural and functional characteristics of amino acid substitutions responsible for phenotypic display of nsSNPs. We also studied the dependence of selective pressure on the structural and functional properties of proteins. We found that in our dataset the selection pressure against deleterious SNPs depends on the molecular function of the protein, although it is insensitive to several other protein features considered. The strongest selective pressure was detected for proteins involved in transcription regulation.
Figures

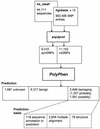
References
-
- Risch N. and Merikangas,K. (1996) The future of genetic studies of complex human diseases. Science, 273, 1516–1517. - PubMed
-
- Risch N.J. (2000) Searching for genetic determinants in the new millennium. Nature, 15, 847–856. - PubMed
-
- Lai E., Riley,J., Purvis,I. and Roses,A. (1998) A 4-Mb high-density single nucleotide polymorphism-based map around human APOE. Genomics, 54, 31–38. - PubMed
-
- Emahazion T., Feuk,L., Jobs,M., Sawyer,S.L., Fredman,D., St Clair,D., Prince,J.A. and Brookes,A.J. (2001) SNP association studies in Alzheimer’s disease highlight problem for complex disease analysis. Trends Genet., 17, 407–413. - PubMed
-
- Schork N.J., Fallin,D. and Lanchbury,J.S. (2000) Single nucleotide polymorphisms and the future of genetic epidemiology. Clin. Genet., 58, 250–264. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

