Suppression of transgene silencing by matrix attachment regions in maize: a dual role for the maize 5' ADH1 matrix attachment region
- PMID: 12215518
- PMCID: PMC150768
- DOI: 10.1105/tpc.004028
Suppression of transgene silencing by matrix attachment regions in maize: a dual role for the maize 5' ADH1 matrix attachment region
Abstract
Matrix attachment regions (MARs) are DNA sequences that bind an internal nuclear network of nonhistone proteins called the nuclear matrix. Thus, they may define discrete gene-containing chromatin loops in vivo. We have studied the effects of flanking transgenes with MARs on transgene expression levels in maize callus and in transformed maize plants. Three MAR elements, two from maize (Adh1 5' MAR and Mha1 5' MAR) and one from yeast (ARS1), had very different effects on transgene expression that bore no relation to their affinity for the nuclear matrix in vitro. In callus, two of the MAR elements (Adh1 5' MAR and ARS1) reduced transgene silencing but had no effect on the variability of expression. In transgenic plants, Adh1 5' MAR had the effect of localizing beta-glucuronidase expression to lateral root initiation sites. A possible model accounting for the function of Adh1 5' MAR is discussed.
Figures

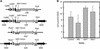




Similar articles
-
Functional analysis of two matrix attachment region (MAR) elements in transgenic maize plants.Transgenic Res. 2003 Apr;12(2):137-54. doi: 10.1023/a:1022908614356. Transgenic Res. 2003. PMID: 12739882
-
Use of matrix attachment regions (MARs) to minimize transgene silencing.Plant Mol Biol. 2000 Jun;43(2-3):361-76. doi: 10.1023/a:1006424621037. Plant Mol Biol. 2000. PMID: 10999416 Review.
-
Elevation of transgene expression level by flanking matrix attachment regions (MAR) is promoter dependent: a study of the interactions of six promoters with the RB7 3' MAR.Transgenic Res. 2003 Feb;12(1):3-12. doi: 10.1023/a:1022194120518. Transgenic Res. 2003. PMID: 12650520
-
The matrix attachment regions (MARs) associated with the Heat Shock Cognate 80 gene ( HSC80) of tomato represent specific regulatory elements.Mol Genet Genomics. 2002 Jan;266(5):891-8. doi: 10.1007/s00438-001-0613-x. Epub 2001 Nov 29. Mol Genet Genomics. 2002. PMID: 11810265
-
[The potential role of nuclear matrix attachment regions (MARs) in regulation of gene expression].Sheng Wu Gong Cheng Xue Bao. 2004 Jan;20(1):6-9. Sheng Wu Gong Cheng Xue Bao. 2004. PMID: 16108480 Review. Chinese.
Cited by
-
Minimizing the unpredictability of transgene expression in plants: the role of genetic insulators.Plant Cell Rep. 2012 Jan;31(1):13-25. doi: 10.1007/s00299-011-1167-y. Epub 2011 Oct 11. Plant Cell Rep. 2012. PMID: 21987122 Review.
-
Functional analysis of the HS185 regulatory element in the rice HSP70 promoter.Mol Biol Rep. 2012 Feb;39(2):1649-57. doi: 10.1007/s11033-011-0904-1. Epub 2011 Jun 3. Mol Biol Rep. 2012. PMID: 21633891
-
Mapping of scaffold/matrix attachment regions in human genome: a data mining exercise.Nucleic Acids Res. 2019 Aug 22;47(14):7247-7261. doi: 10.1093/nar/gkz562. Nucleic Acids Res. 2019. PMID: 31265077 Free PMC article.
-
The presence of a chromatin boundary appears to shield a transgene in tobacco from RNA silencing.Plant Cell. 2003 Sep;15(9):2203-17. doi: 10.1105/tpc.012070. Plant Cell. 2003. PMID: 12953121 Free PMC article.
-
Heritable transgene expression pattern imposed onto maize ubiquitin promoter by maize adh-1 matrix attachment regions: tissue and developmental specificity in maize transgenic plants.Plant Cell Rep. 2004 Jul;22(12):931-8. doi: 10.1007/s00299-004-0779-x. Epub 2004 May 4. Plant Cell Rep. 2004. PMID: 15127223
References
-
- Amati, B.B., and Gasser, S.M. (1988). Chromosomal ARS and CEN elements bind specifically to the yeast nuclear scaffold. Cell 54, 967–978. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

