Histone modifications depict an aberrantly heterochromatinized FMR1 gene in fragile x syndrome
- PMID: 12232854
- PMCID: PMC378545
- DOI: 10.1086/342931
Histone modifications depict an aberrantly heterochromatinized FMR1 gene in fragile x syndrome
Abstract
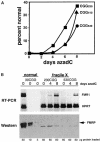
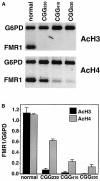
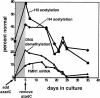
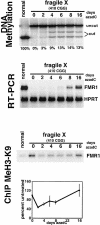
Fragile X syndrome is caused by an expansion of a polymorphic CGG triplet repeat that results in silencing of FMR1 expression. This expansion triggers methylation of FMR1's CpG island, hypoacetylation of associated histones, and chromatin condensation, all characteristics of a transcriptionally inactive gene. Here, we show that there is a graded spectrum of histone H4 acetylation that is proportional to CGG repeat length and that correlates with responsiveness of the gene to DNA demethylation but not with chromatin condensation. We also identify alterations in patient cells of two recently identified histone H3 modifications: methylation of histone H3 at lysine 4 and methylation of histone H3 at lysine 9, which are marks for euchromatin and heterochromatin, respectively. In fragile X cells, there is a decrease in methylation of histone H3 at lysine 4 with a large increase in methylation at lysine 9, a change that is consistent with the model of FMR1's switch from euchromatin to heterochromatin in the disease state. The high level of histone H3 methylation at lysine 9 may account for the failure of H3 to be acetylated after treatment of fragile X cells with inhibitors of histone deacetylases, a treatment that fully restores acetylation to histone H4. Using 5-aza-2'-deoxycytidine, we show that DNA methylation is tightly coupled to the histone modifications associated with euchromatin but not to the heterochromatic mark of methylation of histone H3 at lysine 9, consistent with recent findings that this histone modification may direct DNA methylation. Despite the drug-induced accumulation of mRNA in patient cells to 35% of the wild-type level, FMR1 protein remained undetectable. The identification of intermediates in the heterochromatinization of FMR1 has enabled us to begin to dissect the epigenetics of silencing of a disease-related gene in its natural chromosomal context.
Figures


References
Electronic-Database Information
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for FMR1 [MIM 309550])
References
-
- Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC, Kouzarides T (2001) Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromodomain. Nature 410:120–124 - PubMed
-
- Bestor TH (1998) Methylation meets acetylation. Nature 393:311–312 - PubMed
-
- Boggs BA, Cheung P, Heard E, Spector DL, Chinault AC, Allis CD (2002) Differentially methylated forms of histone H3 show unique association patterns with inactive human X chromosomes. Nat Genet 30:73–76 - PubMed
-
- Chiurazzi P, Pomponi MG, Pietrobono R, Bakker CE, Neri G, Oostra BA (1999) Synergistic effect of histone hyperacetylation and DNA demethylation in the reactivation of the FMR1 gene. Hum Mol Genet 8:2317–2323 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

