Endogenous expression of a high-affinity pseudoknot RNA aptamer suppresses replication of HIV-1
- PMID: 12235384
- PMCID: PMC137107
- DOI: 10.1093/nar/gkf522
Endogenous expression of a high-affinity pseudoknot RNA aptamer suppresses replication of HIV-1
Abstract
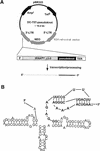
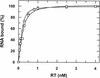
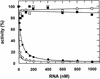
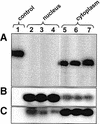
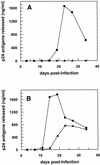
Aptamers, small oligonucleotides derived from an in vitro evolution process called SELEX, are promising therapeutic and diagnostic agents. Although very effective in vitro, only a few examples are available showing their potential in vivo. We have analyzed the effect of a well characterized pseudoknot RNA aptamer selected for tight binding to human immunodeficiency virus (HIV) type 1 reverse transcriptase on HIV replication. Transient intracellular expression of a chimeric RNA consisting of the human initiator tRNA(Met) (tRNA(Meti))/aptamer sequence in human 293T cells showed inhibition of HIV particle release by >75% when the cells were co-transfected with proviral HIV-1 DNA. Subsequent virus production of human T-lymphoid C8166 cells, infected with viral particles derived from co-transfected 293T cells, was again reduced by >75% as compared with the control. As the observed effects are additive, in this model for virus spread, the total reduction of HIV particle formation by transient intracellular expression of the pseudoknot RNA aptamer amounts to >95%. Low-dose HIV infection of human T cells stably expressing the aptamer did not show any virus replication over a period of 35 days. This is the first example of an RNA aptamer selected against a viral enzyme target to show powerful antiviral activity in HIV-1-permissive human T-lymphoid cell lines.
Figures

Similar articles
-
Potent inhibition of human immunodeficiency virus type 1 replication by template analog reverse transcriptase inhibitors derived by SELEX (systematic evolution of ligands by exponential enrichment).J Virol. 2002 Jul;76(13):6545-57. doi: 10.1128/jvi.76.13.6545-6557.2002. J Virol. 2002. PMID: 12050367 Free PMC article.
-
Receptor ligand-facilitated cationic liposome delivery of anti-HIV-1 Rev-binding aptamer and ribozyme DNAs.J Drug Target. 1998;5(4):247-59. doi: 10.3109/10611869808995879. J Drug Target. 1998. PMID: 9713975
-
Inhibition of HIV-1 in CEM cells by a potent TAR decoy.Gene Ther. 1995 Aug;2(6):377-84. Gene Ther. 1995. PMID: 7584112
-
Anti-HIV inhibitors based on nucleic acids: emergence of aptamers as potent antivirals.Curr Drug Targets Infect Disord. 2003 Dec;3(4):383-400. doi: 10.2174/1568005033481060. Curr Drug Targets Infect Disord. 2003. PMID: 14754437 Review.
-
HIV-1 as RNA evolution machine.RNA Biol. 2011 Mar-Apr;8(2):225-9. doi: 10.4161/rna.8.2.14801. Epub 2011 Mar 1. RNA Biol. 2011. PMID: 21358278 Review.
Cited by
-
High-affinity RNA Aptamers Against the HIV-1 Protease Inhibit Both In Vitro Protease Activity and Late Events of Viral Replication.Mol Ther Nucleic Acids. 2015 Feb 17;4(2):e228. doi: 10.1038/mtna.2015.1. Mol Ther Nucleic Acids. 2015. PMID: 25689224 Free PMC article.
-
Potent Inhibition of HIV-1 Reverse Transcriptase and Replication by Nonpseudoknot, "UCAA-motif" RNA Aptamers.Mol Ther Nucleic Acids. 2013 Feb 5;2(2):e71. doi: 10.1038/mtna.2012.62. Mol Ther Nucleic Acids. 2013. PMID: 23385524 Free PMC article.
-
Oligomeric nucleic acids as antivirals.Molecules. 2011 Jan 28;16(2):1271-96. doi: 10.3390/molecules16021271. Molecules. 2011. PMID: 21278679 Free PMC article. Review.
-
Novel bimodular DNA aptamers with guanosine quadruplexes inhibit phylogenetically diverse HIV-1 reverse transcriptases.Nucleic Acids Res. 2008 Dec;36(22):7124-35. doi: 10.1093/nar/gkn891. Epub 2008 Nov 7. Nucleic Acids Res. 2008. PMID: 18996899 Free PMC article.
-
Numerical integration methods and layout improvements in the context of dynamic RNA visualization.BMC Bioinformatics. 2017 May 30;18(1):282. doi: 10.1186/s12859-017-1682-0. BMC Bioinformatics. 2017. PMID: 28558664 Free PMC article.
References
-
- Robertson D.L. and Joyce,G.F. (1990) Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature, 344, 467–468. - PubMed
-
- Tuerk C. and Gold,L. (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science, 249, 505–510. - PubMed
-
- Ellington A.D. and Szostak,J.W. (1990) In vitro selection of RNA molecules that bind specific ligands. Nature, 346, 818–822. - PubMed
-
- Brody E.N. and Gold,L. (2000) Aptamers as therapeutic and diagnostic agents. J. Biotechnol., 74, 5–13. - PubMed
-
- Famulok M., Mayer,G. and Blind,M. (2000) Nucleic acid aptamers-from selection in vitro to applications in vivo. Acc. Chem. Res., 33, 591–599. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

