EcoRII: a restriction enzyme evolving recombination functions?
- PMID: 12356742
- PMCID: PMC129036
- DOI: 10.1093/emboj/cdf514
EcoRII: a restriction enzyme evolving recombination functions?
Abstract
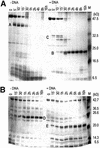
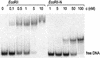
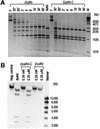
The restriction endonuclease EcoRII requires the cooperative interaction with two copies of the sequence 5'CCWGG for DNA cleavage. We found by limited proteolysis that EcoRII has a two-domain structure that enables this particular mode of protein-DNA interaction. The C-terminal domain is a new restriction endonuclease, EcoRII-C. In contrast to the wild-type enzyme, EcoRII-C cleaves DNA specifically at single 5'CCWGG sites. Moreover, substrates containing two or more cooperative 5'CCWGG sites are cleaved much more efficiently by EcoRII-C than by EcoRII. The N-terminal domain binds DNA specifically and attenuates the activity of EcoRII by making the enzyme dependent on a second 5'CCWGG site. Therefore, we suggest that a precursor EcoRII endonuclease acquired an additional DNA-binding domain to enable the interaction with two 5'CCWGG sites. The current EcoRII molecule could be an evolutionary intermediate between a site-specific endonuclease and a protein that functions specifically with two DNA sites such as recombinases and transposases. The combination of these functions may enable EcoRII to accomplish its own propagation similarly to transposons.
Figures


Similar articles
-
Crystal structure of type IIE restriction endonuclease EcoRII reveals an autoinhibition mechanism by a novel effector-binding fold.J Mol Biol. 2004 Jan 2;335(1):307-19. doi: 10.1016/j.jmb.2003.10.030. J Mol Biol. 2004. PMID: 14659759
-
Mechanism of the interaction of EcoRII restriction endonuclease with two recognition sites. Probing of modified DNA duplexes as activators of the enzyme.Eur J Biochem. 1992 Sep 15;208(3):617-22. doi: 10.1111/j.1432-1033.1992.tb17226.x. Eur J Biochem. 1992. PMID: 1396668
-
Two subunits of EcoRII restriction endonuclease interact with two DNA recognition sites.Biochem Biophys Res Commun. 1994 Feb 15;198(3):885-90. doi: 10.1006/bbrc.1994.1126. Biochem Biophys Res Commun. 1994. PMID: 8117292
-
Cooperative binding properties of restriction endonuclease EcoRII with DNA recognition sites.J Biol Chem. 1998 Apr 3;273(14):8294-300. doi: 10.1074/jbc.273.14.8294. J Biol Chem. 1998. PMID: 9525936
-
Molecular analysis of restriction endonuclease EcoRII from Escherichia coli reveals precise regulation of its enzymatic activity by autoinhibition.Mol Microbiol. 2009 May;72(4):1011-21. doi: 10.1111/j.1365-2958.2009.06702.x. Epub 2009 Apr 21. Mol Microbiol. 2009. PMID: 19400796
Cited by
-
Direct monitoring of allosteric recognition of type IIE restriction endonuclease EcoRII.J Biol Chem. 2008 May 30;283(22):15023-30. doi: 10.1074/jbc.M800334200. Epub 2008 Mar 26. J Biol Chem. 2008. PMID: 18367450 Free PMC article.
-
Structure of the metal-independent restriction enzyme BfiI reveals fusion of a specific DNA-binding domain with a nonspecific nuclease.Proc Natl Acad Sci U S A. 2005 Nov 1;102(44):15797-802. doi: 10.1073/pnas.0507949102. Epub 2005 Oct 24. Proc Natl Acad Sci U S A. 2005. PMID: 16247004 Free PMC article.
-
A homology model of restriction endonuclease SfiI in complex with DNA.BMC Struct Biol. 2005 Jan 24;5:2. doi: 10.1186/1472-6807-5-2. BMC Struct Biol. 2005. PMID: 15667656 Free PMC article.
-
Bacteriophage T4 endonuclease II, a promiscuous GIY-YIG nuclease, binds as a tetramer to two DNA substrates.Nucleic Acids Res. 2009 Oct;37(18):6174-83. doi: 10.1093/nar/gkp652. Epub 2009 Aug 7. Nucleic Acids Res. 2009. PMID: 19666720 Free PMC article.
-
LraI from Lactococcus raffinolactis BGTRK10-1, an Isoschizomer of EcoRI, Exhibits Ion Concentration-Dependent Specific Star Activity.Biomed Res Int. 2018 Jan 29;2018:5657085. doi: 10.1155/2018/5657085. eCollection 2018. Biomed Res Int. 2018. PMID: 29789800 Free PMC article.
References
-
- Arber W. (2000) Genetic variation: molecular mechanisms and impact on microbial evolution. FEMS Microbiol. Rev., 24, 1–7. - PubMed
-
- Babbitt P.C. and Gerlt,J.A. (2000) New functions from old scaffolds: how nature reengineers enzymes for new functions. Adv. Protein Chem., 55, 1–28. - PubMed
-
- Behlke J., Ristau,O. and Schonfeld,H.J. (1997) Nucleotide-dependent complex formation between the Escherichia coli chaperonins GroEL and GroES studied under equilibrium conditions. Biochemistry, 36, 5149–5156. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

