Extracellular protons regulate the extracellular cation selectivity of the sodium pump
- PMID: 12356852
- PMCID: PMC2229530
- DOI: 10.1085/jgp.20028573
Extracellular protons regulate the extracellular cation selectivity of the sodium pump
Abstract
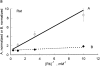
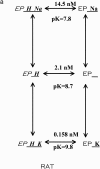
The effects of 0.3-10 nM extracellular protons (pH 9.5-8.0) on ouabain-sensitive rubidium influx were determined in 4,4'-diisocyanostilbene-2, 2'-disulfonate (DIDS)-treated human and rat erythrocytes. This treatment clamps the intracellular H. We found that rubidium binds much better to the protonated pump than the unprotonated pump; 13-fold better in rat and 34-fold better in human erythrocytes. This clearly shows that protons are not competing with rubidium in this proton concentration range. Bretylium and tetrapropylammonium also bind much better to the protonated pump than the unprotonated pump in human erythrocytes and in this sense they are potassium-like ions. In contrast, guanidinium and sodium bind about equally well to protonated and unprotonated pump in human red cells. In rat red cells, protons actually make sodium bind less well (about sevenfold). Thus, protons have substantially different effects on the binding of rubidium and sodium. The effect of protons on ouabain binding in rat red cells was intermediate between the effects of protons on rubidium binding and on sodium binding. Remarkably, all four cationic inhibitors (bretylium, guanidinium, sodium, and tetrapropylammonium) had similar apparent inhibitory constants for the unprotonated pump ( approximately 5-10 mM). The K(d) for proton binding to the human pump, with the empty transport site facing extracellularly is 13 nM, whereas the extracellular transport site loaded with sodium is 9.5 nM, and with rubidium is 0.38 nM. In rat red cells there is also a substantial difference in the K(d) for proton binding to the sodium-loaded pump (14.5 nM) and the rubidium-loaded pump (0.158 nM). These data suggest that important rearrangements occur at the extracellular pump surface as the pump moves between conformations in which the outward facing transport site has sodium bound, is empty, or has rubidium bound and that guanidinium is sodium-like and bretylium and tetrapropylammonium are rubidium-like.
Figures














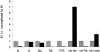

Similar articles
-
Red blood cell Na pump: Insights from species differences.Blood Cells Mol Dis. 2009 May-Jun;42(3):192-200. doi: 10.1016/j.bcmd.2009.01.011. Epub 2009 Mar 5. Blood Cells Mol Dis. 2009. PMID: 19268612 Free PMC article. Review.
-
Palytoxin induces an increase in the cation conductance of red cells.J Gen Physiol. 1991 Nov;98(5):969-85. doi: 10.1085/jgp.98.5.969. J Gen Physiol. 1991. PMID: 1684984 Free PMC article.
-
Bretylium, an organic quaternary amine, inhibits the Na,K-ATPase by binding to the extracellular K-site.Blood Cells Mol Dis. 2004 May-Jun;32(3):394-400. doi: 10.1016/j.bcmd.2004.01.013. Blood Cells Mol Dis. 2004. PMID: 15121098
-
Evidence for tryptophan residues in the cation transport path of the Na(+),K(+)-ATPase.Biochemistry. 2003 Sep 2;42(34):10212-22. doi: 10.1021/bi0342721. Biochemistry. 2003. PMID: 12939149
-
The sodium pump becomes a family.Trends Neurosci. 1988 Jul;11(7):325-8. doi: 10.1016/0166-2236(88)90096-3. Trends Neurosci. 1988. PMID: 2465638 Review. No abstract available.
Cited by
-
Molecular simulations and free-energy calculations suggest conformation-dependent anion binding to a cytoplasmic site as a mechanism for Na+/K+-ATPase ion selectivity.J Biol Chem. 2017 Jul 28;292(30):12412-12423. doi: 10.1074/jbc.M117.779090. Epub 2017 Jun 6. J Biol Chem. 2017. PMID: 28588025 Free PMC article.
-
Distinct pH dependencies of Na+/K+ selectivity at the two faces of Na,K-ATPase.J Biol Chem. 2018 Feb 9;293(6):2195-2205. doi: 10.1074/jbc.RA117.000700. Epub 2017 Dec 15. J Biol Chem. 2018. PMID: 29247005 Free PMC article.
-
Protonation of key acidic residues is critical for the K⁺-selectivity of the Na/K pump.Nat Struct Mol Biol. 2011 Sep 11;18(10):1159-63. doi: 10.1038/nsmb.2113. Nat Struct Mol Biol. 2011. PMID: 21909093 Free PMC article.
-
Red blood cell Na pump: Insights from species differences.Blood Cells Mol Dis. 2009 May-Jun;42(3):192-200. doi: 10.1016/j.bcmd.2009.01.011. Epub 2009 Mar 5. Blood Cells Mol Dis. 2009. PMID: 19268612 Free PMC article. Review.
References
-
- Apell, H.J., and S.J. Karlish. 2001. Functional properties of Na,K-ATPase, and their structural implications, as detected with biophysical techniques. J. Membr. Biol. 180:1–9. - PubMed
-
- Blostein, R. 1999. Jeanne Mannery Fisher Memorial Lecture 1998. Structure-function studies of the sodium pump. Biochem. Cell Biol. 77:1–10. - PubMed
-
- Breitweiser, G.E., A.A. Altamirano, and J.M. Russell. 1987. Effects of pH changes on sodium pump fluxes in squid giant axon. Am. J. Physiol. Cell Physiol. 253:C547–C554. - PubMed
-
- Cornelius, F., and J.C. Skou. 1985. Na+-Na+ exchange mediated by (Na+ + K+)-ATPase reconstituted into liposomes. Evaluation of pump stoichiometry and response to ATP and ADP. Biochim. Biophys. Acta. 818:211–221. - PubMed

