Inferring domain-domain interactions from protein-protein interactions
- PMID: 12368246
- PMCID: PMC187530
- DOI: 10.1101/gr.153002
Inferring domain-domain interactions from protein-protein interactions
Abstract
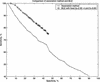
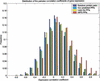
The interaction between proteins is one of the most important features of protein functions. Behind protein-protein interactions there are protein domains interacting physically with one another to perform the necessary functions. Therefore, understanding protein interactions at the domain level gives a global view of the protein interaction network, and possibly of protein functions. Two research groups used yeast two-hybrid assays to generate 5719 interactions between proteins of the yeast Saccharomyces cerevisiae. This allows us to study the large-scale conserved patterns of interactions between protein domains. Using evolutionarily conserved domains defined in a protein-domain database called PFAM (http://PFAM.wustl.edu), we apply a Maximum Likelihood Estimation method to infer interacting domains that are consistent with the observed protein-protein interactions. We estimate the probabilities of interactions between every pair of domains and measure the accuracies of our predictions at the protein level. Using the inferred domain-domain interactions, we predict interactions between proteins. Our predicted protein-protein interactions have a significant overlap with the protein-protein interactions (MIPS: http://mips.gfs.de) obtained by methods other than the two-hybrid assays. The mean correlation coefficient of the gene expression profiles for our predicted interaction pairs is significantly higher than that for random pairs. Our method has shown robustness in analyzing incomplete data sets and dealing with various experimental errors. We found several novel protein-protein interactions such as RPS0A interacting with APG17 and TAF40 interacting with SPT3, which are consistent with the functions of the proteins.
Figures

References
-
- Bock JR, Gough DA. Predicting protein–protein interactions from primary structure. Bioinformatics. 2001;17:455–460. - PubMed
-
- Dempster AP, Laid NM, Rubin DB. Maximum likelihood from incomplete data via the EM algorithm (with discussion) J Roy Statist Soc Ser B. 1977;39:1–38.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
