Differential expression of two distinct phenylalanine ammonia-lyase genes in condensed tannin-accumulating and lignifying cells of quaking aspen
- PMID: 12376645
- PMCID: PMC166607
- DOI: 10.1104/pp.006262
Differential expression of two distinct phenylalanine ammonia-lyase genes in condensed tannin-accumulating and lignifying cells of quaking aspen
Abstract
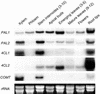
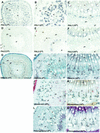
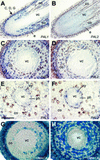
Lignins, along with condensed tannins (CTs) and salicylate-derived phenolic glycosides, constitute potentially large phenylpropanoid carbon sinks in tissues of quaking aspen (Populus tremuloides Michx.). Metabolic commitment to each of these sinks varies during development and adaptation, and depends on L-phenylalanine ammonia-lyase (PAL), an enzyme catalyzing the deamination of L-phenylalanine to initiate phenylpropanoid metabolism. In Populus spp., PAL is encoded by multiple genes whose expression has been associated with lignification in primary and secondary tissues. We now report cloning two differentially expressed PAL cDNAs that exhibit distinct spatial associations with CT and lignin biosynthesis in developing shoot and root tissues of aspen. PtPAL1 was expressed in certain CT-accumulating, non-lignifying cells of stems, leaves, and roots, and the pattern of PtPAL1 expression varied coordinately with that of CT accumulation along the primary to secondary growth transition in stems. PtPAL2 was expressed in heavily lignified structural cells of shoots, but was also expressed in non-lignifying cells of root tips. Evidence of a role for Pt4CL2, encoding 4-coumarate:coenzyme A ligase, in determining CT sink strength was gained from cellular co-expression analysis with PAL1 and CTs, and from experiments in which leaf wounding increased PAL1 and 4CL2 expression as well as the relative allocation of carbon to CT with respect to phenolic glycoside, the dominant phenolic sink in aspen leaves. Leaf wounding also increased PAL2 and lignin pathway gene expression, but to a smaller extent. The absence of PAL2 in most CT-accumulating cells provides in situ support for the idea that PAL isoforms function in specific metabolic milieus.
Figures


References
-
- Allwood EG, Davies DR, Gerrish C, Ellis BE, Bolwell GP. Phosphorylation of phenylalanine ammonia-lyase: evidence for a novel protein kinase and identification of the phosphorylated residue. FEBS Lett. 1999;457:47–52. - PubMed
-
- Appert C, Logemann E, Hahlbrock K, Schmid J, Amrhein N. Structural and catalytic properties of the four phenylalanine ammonia-lyase isoenzymes from parsley (Petroselinum crispum Nym.) Eur J Biochem. 1994;225:491–499. - PubMed
-
- Bolwell GP. A role for phosphorylation in the down-regulation of phenylalanine ammonia-lyase in suspension-cultured cells of French bean. Phytochemistry. 1992;31:4081–4086.
-
- Bolwell GP, Cramer CL, Lamb CJ, Schuch W, Dixon RA. l- Phenylalanine ammonia-lyase from Phaseolus vulgaris: modulation of the levels of active enzyme by trans-cinnamic acid. Planta. 1986;169:97–107. - PubMed
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

