A method for prediction of the locations of linker regions within large multifunctional proteins, and application to a type I polyketide synthase
- PMID: 12381311
- PMCID: PMC3400148
- DOI: 10.1016/s0022-2836(02)00972-5
A method for prediction of the locations of linker regions within large multifunctional proteins, and application to a type I polyketide synthase
Abstract
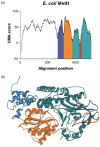
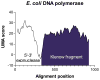
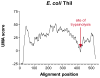
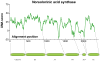
Multifunctional proteins often appear to result from fusion of smaller proteins and in such cases typically can be separated into their ancestral components simply by cleaving the linker regions that separate the domains. Though possibly guided by sequence alignment, structural evidence, or light proteolysis, determination of the locations of linker regions remains empirical. We have developed an algorithm, named UMA, to predict the locations of linker regions in multifunctional proteins by quantification of the conservation of several properties within protein families, and the results agree well with structurally characterized proteins. This technique has been applied to a family of fungal type I iterative polyketide synthases (PKS), allowing prediction of the locations of all of the standard PKS domains, as well as two previously unidentified domains. Using these predictions, we report the cloning of the first fragment from the PKS norsolorinic acid synthase, responsible for biosynthesis of the first isolatable intermediate in aflatoxin production. The expression, light proteolysis and catalytic abilities of this acyl carrier protein-thioesterase didomain are discussed.
Figures




References
-
- Cohen GB, Ren R, Baltimore D. Modular binding domains in signal transduction proteins. Cell. 1995;80:237–248. - PubMed
-
- Bandarian V, Pattridge KA, Lennon BW, Huddler DP, Matthews RG, Ludwig ML. Domain alternation switches B(12)-dependent methionine synthase to the activation conformation. Nature Struct Biol. 2002;9:53–56. - PubMed
-
- Thoden JB, Miran SG, Phillips JC, Howard AJ, Raushel FM, Holden HM. Carbamoyl phosphate synthetase: caught in the act of glutamine hydrolysis. Biochemistry. 1998;37:8825–8831. - PubMed
-
- Staunton J, Weissman KJ. Polyketide biosynthesis: a millennium review. Nature Prod Rep. 2001;18:380–416. - PubMed
-
- Marahiel MA, Stachelhaus T, Mootz HD. Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem Rev. 1997;97:2651–2674. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

