Determinants of Swe1p degradation in Saccharomyces cerevisiae
- PMID: 12388757
- PMCID: PMC129966
- DOI: 10.1091/mbc.e02-05-0283
Determinants of Swe1p degradation in Saccharomyces cerevisiae
Abstract
Swe1p, the sole Wee1-family kinase in Saccharomyces cerevisiae, is synthesized during late G1 and is then degraded as cells proceed through the cell cycle. However, Swe1p degradation is halted by the morphogenesis checkpoint, which responds to insults that perturb bud formation. The Swe1p stabilization promotes cell cycle arrest through Swe1p-mediated inhibitory phosphorylation of Cdc28p until the cells can recover from the perturbation and resume bud formation. Swe1p degradation involves the relocalization of Swe1p from the nucleus to the mother-bud neck, and neck targeting requires the Swe1p-interacting protein Hsl7p. In addition, Swe1p degradation is stimulated by its substrate, cyclin/Cdc28p, and Swe1p is thought to be a target of the ubiquitin ligase SCF(Met30) acting with the ubiquitin-conjugating enzyme Cdc34p. The basis for regulation of Swe1p degradation by the morphogenesis checkpoint remains unclear, and in order to elucidate that regulation we have dissected the Swe1p degradation pathway in more detail, yielding several novel findings. First, we show here that Met30p (and by implication SCF(Met30)) is not, in fact, required for Swe1p degradation. Second, cyclin/Cdc28p does not influence Swe1p neck targeting, but can directly phosphorylate Swe1p, suggesting that it acts downstream of neck targeting in the Swe1p degradation pathway. Third, a screen for functional but nondegradable mutants of SWE1 identified two small regions of Swe1p that are key to its degradation. One of these regions mediates interaction of Swe1p with Hsl7p, showing that the Swe1p-Hsl7p interaction is critical for Swe1p neck targeting and degradation. The other region did not appear to affect interactions with known Swe1p regulators, suggesting that other as-yet-unknown regulators exist.
Figures







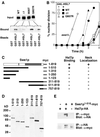


Similar articles
-
The morphogenesis checkpoint in Saccharomyces cerevisiae: cell cycle control of Swe1p degradation by Hsl1p and Hsl7p.Mol Cell Biol. 1999 Oct;19(10):6929-39. doi: 10.1128/MCB.19.10.6929. Mol Cell Biol. 1999. PMID: 10490630 Free PMC article.
-
Roles of Hsl1p and Hsl7p in Swe1p degradation: beyond septin tethering.Eukaryot Cell. 2012 Dec;11(12):1496-502. doi: 10.1128/EC.00196-12. Epub 2012 Oct 5. Eukaryot Cell. 2012. PMID: 23042131 Free PMC article.
-
Septin-dependent assembly of a cell cycle-regulatory module in Saccharomyces cerevisiae.Mol Cell Biol. 2000 Jun;20(11):4049-61. doi: 10.1128/MCB.20.11.4049-4061.2000. Mol Cell Biol. 2000. PMID: 10805747 Free PMC article.
-
Eavesdropping on the cytoskeleton: progress and controversy in the yeast morphogenesis checkpoint.Curr Opin Microbiol. 2006 Dec;9(6):540-6. doi: 10.1016/j.mib.2006.10.004. Epub 2006 Oct 19. Curr Opin Microbiol. 2006. PMID: 17055334 Review.
-
The morphogenesis checkpoint: how yeast cells watch their figures.Curr Opin Cell Biol. 2003 Dec;15(6):648-53. doi: 10.1016/j.ceb.2003.09.001. Curr Opin Cell Biol. 2003. PMID: 14644188 Review.
Cited by
-
Kinetic partitioning during de novo septin filament assembly creates a critical G1 "window of opportunity" for mutant septin function.Cell Cycle. 2016 Sep 16;15(18):2441-53. doi: 10.1080/15384101.2016.1196304. Epub 2016 Jul 11. Cell Cycle. 2016. PMID: 27398993 Free PMC article.
-
The yeast ubiquitin ligase SCFMet30: connecting environmental and intracellular conditions to cell division.Cell Div. 2006 Aug 8;1:16. doi: 10.1186/1747-1028-1-16. Cell Div. 2006. PMID: 16895602 Free PMC article.
-
Prolonged cyclin-dependent kinase inhibition results in septin perturbations during return to growth and mitosis.J Cell Biol. 2018 Jul 2;217(7):2429-2443. doi: 10.1083/jcb.201708153. Epub 2018 May 9. J Cell Biol. 2018. PMID: 29743192 Free PMC article.
-
ELM1 is required for multidrug resistance in Saccharomyces cerevisiae.Genetics. 2006 Aug;173(4):1919-37. doi: 10.1534/genetics.106.057596. Epub 2006 Jun 4. Genetics. 2006. PMID: 16751665 Free PMC article.
-
Cdc34 C-terminal tail phosphorylation regulates Skp1/cullin/F-box (SCF)-mediated ubiquitination and cell cycle progression.Biochem J. 2007 Aug 1;405(3):569-81. doi: 10.1042/BJ20061812. Biochem J. 2007. PMID: 17461777 Free PMC article.
References
-
- Bao S, et al. The highly conserved protein methyltransferase, Skb1, is a mediator of hyperosmotic stress response in the fission yeast Schizosaccharomyces pombe. J Biol Chem. 2001;276:14549–14552. - PubMed
-
- Bishop AC, et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature. 2000;407:395–401. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

