Regulation of the pollen-specific actin-depolymerizing factor LlADF1
- PMID: 12417710
- PMCID: PMC152736
- DOI: 10.1105/tpc.005363
Regulation of the pollen-specific actin-depolymerizing factor LlADF1
Abstract
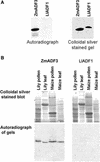
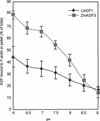
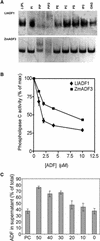
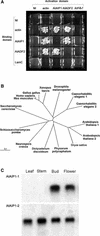
Pollen tube growth is dependent on a dynamic actin cytoskeleton, suggesting that actin-regulating proteins are involved. We have examined the regulation of the lily pollen-specific actin-depolymerizing factor (ADF) LlADF1. Its actin binding and depolymerizing activity is pH sensitive, inhibited by certain phosphoinositides, but not controlled by phosphorylation. Compared with its F-actin binding properties, its low activity in depolymerization assays has been used to explain why pollen ADF decorates F-actin in pollen grains. This low activity is incompatible with a role in increasing actin dynamics necessary to promote pollen tube growth. We have identified a plant homolog of actin-interacting protein, AIP1, which enhances the depolymerization of F-actin in the presence of LlADF1 by approximately 60%. Both pollen ADF and pollen AIP1 bind F-actin in pollen grains but are mainly cytoplasmic in pollen tubes. Our results suggest that together these proteins remodel actin filaments as pollen grains enter and exit dormancy.
Figures


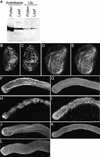
References
-
- Aizawa, H., Katadae, M., Maruya, M., Sameshima, M., Murakami-Murofushi, K., and Yahara, I. (1999). Hyperosmotic stress-induced reorganization of actin bundles in Dictyostelium cells over-expressing cofilin. Genes Cells 4, 311–324. - PubMed
-
- Allwood, E.G., Smertenko, A.P., and Hussey, P.J. (2001). Phosphorylation of plant actin-depolymerising factor by calmodulin-like domain protein kinase. FEBS Lett. 499, 97–100. - PubMed
-
- Amberg, D.C., Basart, E., and Botstein, D. (1995). Defining protein interactions with yeast actin in-vivo. Nat. Struct. Biol. 2, 28–35. - PubMed
-
- Arber, S., Barbayannis, F.A., Hanser, H., Schneider, C., Stanyon, C.A., Bernard, O., and Caroni, P. (1998). Regulation of actin dynamics through phosphorylation of cofilin by LIM-kinase. Nature 393, 805–809. - PubMed
-
- Ausubel, F.M., Brent, R., Kingston, R.E., Moore, D.D., Seidman, J.G., Smith, J.A., and Struhl, K. (1992). Current Protocols in Molecular Biology. (New York: Green Publishing Associates and Wiley Interscience).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

