Programmed translational frameshift in the bacteriophage P2 FETUD tail gene operon
- PMID: 12426340
- PMCID: PMC135442
- DOI: 10.1128/JB.184.23.6522-6531.2002
Programmed translational frameshift in the bacteriophage P2 FETUD tail gene operon
Abstract
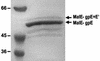
The major structural components of the P2 contractile tail are encoded in the FETUD tail gene operon. The sequences of genes F(I) and F(II), encoding the major tail sheath and tail tube proteins, have been reported previously (L. M. Temple, S. L. Forsburg, R. Calendar, and G. E. Christie, Virology 181:353-358, 1991). Sequence analysis of the remainder of this operon and the locations of amber mutations Eam30, Tam5, Tam64, Tam215, Uam25, Uam77, Uam92, and Dam6 and missense mutation Ets55 identified the coding regions for genes E, T, U, and D, completing the sequence determination of the P2 genome. Inspection of the DNA sequence revealed a new open reading frame overlapping the end of the essential tail gene E. Lack of an apparent translation initiation site and identification of a putative sequence for a programmed translational frameshift within the E gene suggested that this new reading frame (E') might be translated as an extension of gene E, following a -1 translational frameshift. Complementation analysis demonstrated that E' was essential for P2 lytic growth. Analysis of fusion polypeptides verified that this reading frame was translated as a -1 frameshift extension of gpE, with a frequency of approximately 10%. The arrangement of these two genes within the tail gene cluster of phage P2 and their coupling via a translational frameshift appears to be conserved among P2-related phages. This arrangement shows a striking parallel to the organization in the tail gene cluster of phage lambda, despite a lack of amino acid sequence similarity between the tail gene products of these phage families.
Figures

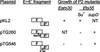


References
-
- Abuladze, N. K., M. Gingery, J. Tsai, and F. A. Eiserling. 1994. Tail length determination in bacteriophage T4. Virology 199:301-310. - PubMed
-
- Bertani, L. E. 1957. The effect of the inhibition of protein synthesis on the establishment of lysogeny. Virology 4:53-71. - PubMed
-
- Bertani, L. E., and E. W. Six. 1988. The P2-like phages and their parasite, P4, p. 73-143. In R. Calendar (ed.), The bacteriophages, vol. 2. Plenum Publishing Corp., New York, N.Y.
-
- Birkeland, N. K., and B. H. Lindqvist. 1986. Coliphage P2 late control gene ogr: DNA sequence and product identification. J. Mol. Biol. 188:487-490. - PubMed
-
- Brosius, J. 1984. Plasmid vectors for the selection of promoters. Gene 27:151-160. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

