Male mouse recombination maps for each autosome identified by chromosome painting
- PMID: 12432495
- PMCID: PMC517487
- DOI: 10.1086/344714
Male mouse recombination maps for each autosome identified by chromosome painting
Abstract
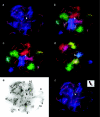
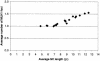
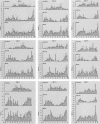
Linkage maps constructed from genetic analysis of gene order and crossover frequency provide few clues to the basis of genomewide distribution of meiotic recombination, such as chromosome structure, that influences meiotic recombination. To bridge this gap, we have generated the first cytological recombination map that identifies individual autosomes in the male mouse. We prepared meiotic chromosome (synaptonemal complex [SC]) spreads from 110 mouse spermatocytes, identified each autosome by multicolor fluorescence in situ hybridization of chromosome-specific DNA libraries, and mapped >2,000 sites of recombination along individual autosomes, using immunolocalization of MLH1, a mismatch repair protein that marks crossover sites. We show that SC length is strongly correlated with crossover frequency and distribution. Although the length of most SCs corresponds to that predicted from their mitotic chromosome length rank, several SCs are longer or shorter than expected, with corresponding increases and decreases in MLH1 frequency. Although all bivalents share certain general recombination features, such as few crossovers near the centromeres and a high rate of distal recombination, individual bivalents have unique patterns of crossover distribution along their length. In addition to SC length, other, as-yet-unidentified, factors influence crossover distribution leading to hot regions on individual chromosomes, with recombination frequencies as much as six times higher than average, as well as cold spots with no recombination. By reprobing the SC spreads with genetically mapped BACs, we demonstrate a robust strategy for integrating genetic linkage and physical contig maps with mitotic and meiotic chromosome structure.
Figures

Similar articles
-
Inter-sex variation in synaptonemal complex lengths largely determine the different recombination rates in male and female germ cells.Cytogenet Genome Res. 2004;107(3-4):208-15. doi: 10.1159/000080599. Cytogenet Genome Res. 2004. PMID: 15467366
-
Discontinuities and unsynapsed regions in meiotic chromosomes have a cis effect on meiotic recombination patterns in normal human males.Hum Mol Genet. 2005 Oct 15;14(20):3013-8. doi: 10.1093/hmg/ddi332. Epub 2005 Sep 9. Hum Mol Genet. 2005. PMID: 16155114
-
Discontinuities and unsynapsed regions in meiotic chromosomes have a trans effect on meiotic recombination of some chromosomes in human males.Cytogenet Genome Res. 2007;119(1-2):27-32. doi: 10.1159/000109615. Epub 2007 Dec 14. Cytogenet Genome Res. 2007. PMID: 18160778
-
Cytological analysis of interference in mouse meiosis.Methods Mol Biol. 2009;558:355-82. doi: 10.1007/978-1-60761-103-5_21. Methods Mol Biol. 2009. PMID: 19685335 Review.
-
[Homologs of MutS and MutL during mammalian meiosis].Med Sci (Paris). 2003 Jan;19(1):85-91. doi: 10.1051/medsci/200319185. Med Sci (Paris). 2003. PMID: 12836196 Review. French.
Cited by
-
A novel function for CDK2 activity at meiotic crossover sites.PLoS Biol. 2020 Oct 19;18(10):e3000903. doi: 10.1371/journal.pbio.3000903. eCollection 2020 Oct. PLoS Biol. 2020. PMID: 33075054 Free PMC article.
-
The Drosophila meiotic mutant mei-352 is an allele of klp3A and reveals a role for a kinesin-like protein in crossover distribution.Genetics. 2005 Aug;170(4):1797-807. doi: 10.1534/genetics.105.041194. Epub 2005 Jun 18. Genetics. 2005. PMID: 15965253 Free PMC article.
-
Hot regions of noninterfering crossovers coexist with a nonuniformly interfering pathway in Arabidopsis thaliana.Genetics. 2013 Nov;195(3):769-79. doi: 10.1534/genetics.113.155549. Epub 2013 Sep 11. Genetics. 2013. PMID: 24026099 Free PMC article.
-
The Genomic Landscape of Crossover Interference in the Desert Tree Populus euphratica.Front Genet. 2019 May 15;10:440. doi: 10.3389/fgene.2019.00440. eCollection 2019. Front Genet. 2019. PMID: 31156703 Free PMC article.
-
MLH1 focus mapping in the guinea fowl (Numida meleagris) give insights into the crossover landscapes in birds.PLoS One. 2020 Oct 5;15(10):e0240245. doi: 10.1371/journal.pone.0240245. eCollection 2020. PLoS One. 2020. PMID: 33017431 Free PMC article.
References
Electronic-Database Information
-
- Genetic and Physical Maps of the Mouse Chromosome, http://www-genome.wi.mit.edu/cgi-bin/mouse/index/ (for BAC clones)
-
- Genome Sequencing, National Center for Biotechnology Information, http://www.ncbi.nlm.nih.gov/genome/seq/
-
- Roswell Park Mouse Screening Project, http://genomics.roswellpark.org/mouse/overview.html (for BAC clones)
References
-
- Badge RM, Yardley J, Jeffreys AJ, Armour JAL (2000) Crossover breakpoint mapping identifies a subtelomeric hotspot for male meiotic recombination. Hum Mol Genet 9:1239–1244 - PubMed
-
- Baker SM, Plug AW, Prolla TA, Bronner CE, Harris AC, Yao X, Christie D-M, Monell C, Arnheim N, Bradley A, Ashley T, Liskay RM (1996) Involvement of mouse Mlh1 in DNA mismatch repair and meiotic crossing over. Nat Genet 13:336–342 - PubMed
-
- Barlow AL, Hultén MA (1998) Crossing over analysis at pachytene in man. Eur J Hum Genet 6:350–358 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

